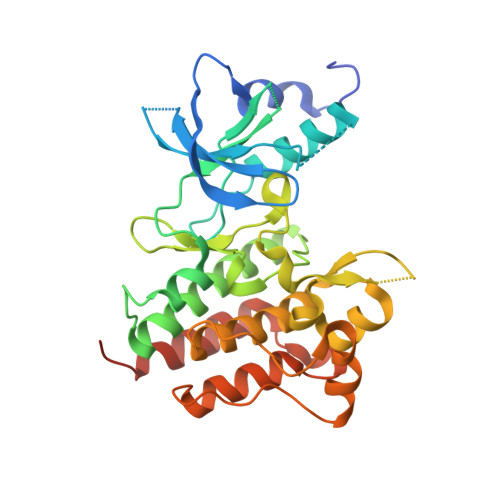
Discovery and Optimization of Triazolopyridazines as Potent and Selective Inhibitors of the c-Met Kinase.
Albrecht, B.K., Harmange, J.C., Bauer, D., Berry, L., Bode, C., Boezio, A.A., Chen, A., Choquette, D., Dussault, I., Fridrich, C., Hirai, S., Hoffman, D., Larrow, J.F., Kaplan-Lefko, P., Lin, J., Lohman, J., Long, A.M., Moriguchi, J., O'Connor, A., Potashman, M.H., Reese, M., Rex, K., Siegmund, A., Shah, K., Shimanovich, R., Springer, S.K., Teffera, Y., Yang, Y., Zhang, Y., Bellon, S.F.(2008) J Med Chem 51: 2879-2882
- PubMed: 18426196 Search on PubMed
- DOI: https://doi.org/10.1021/jm800043g
- Primary Citation Related Structures:
3CCN, 3CD8 - PubMed Abstract:
Tumorigenesis is a multistep process in which oncogenes play a key role in tumor formation, growth, and maintenance. MET was discovered as an oncogene that is activated by its ligand, hepatocyte growth factor. Deregulated signaling in the c-Met pathway has been observed in multiple tumor types. Herein we report the discovery of potent and selective triazolopyridazine small molecules that inhibit c-Met activity.
- Amgen Inc., Cambridge, MA 02139, USA. brian.albrecht@amgen.com
Organizational Affiliation: