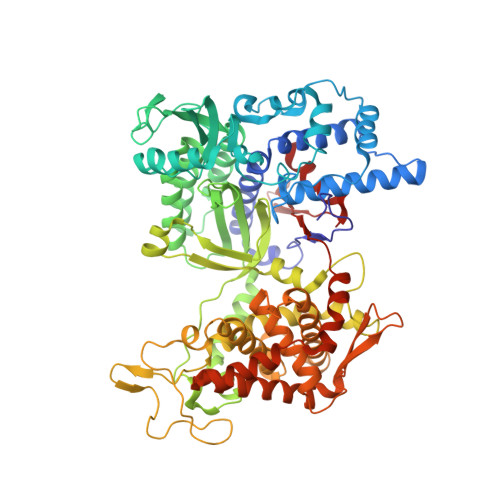
The first structure of dipeptidyl-peptidase III provides insight into the catalytic mechanism and mode of substrate binding.
Baral, P.K., Jajcanin-Jozic, N., Deller, S., Macheroux, P., Abramic, M., Gruber, K.(2008) J Biological Chem 283: 22316-22324
- PubMed: 18550518 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M803522200
- Primary Citation Related Structures:
3CSK - PubMed Abstract:
Dipeptidyl-peptidases III (DPP III) are zinc-dependent enzymes that specifically cleave the first two amino acids from the N terminus of different length peptides. In mammals, DPP III is associated with important physiological functions and is a potential biomarker for certain types of cancer. Here, we present the 1.95-A crystal structure of yeast DPP III representing the prototype for the M49 family of metallopeptidases. It shows a novel fold with two domains forming a wide cleft containing the catalytic metal ion. DPP III exhibits no overall similarity to other metallopeptidases, such as thermolysin and neprilysin, but zinc coordination and catalytically important residues are structurally conserved. Substrate recognition is accomplished by a binding site for the N terminus of the peptide at an appropriate distance from the metal center and by a series of conserved arginine residues anchoring the C termini of different length substrates.
- Institute of Molecular Biosciences, University of Graz, A-8010 Graz, Austria.
Organizational Affiliation: