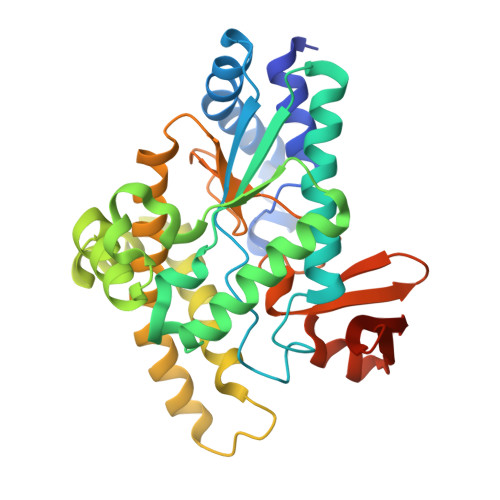
Three-dimensional structures of Pseudomonas aeruginosa PvcA and PvcB, two proteins involved in the synthesis of 2-isocyano-6,7-dihydroxycoumarin.
Drake, E.J., Gulick, A.M.(2008) J Mol Biology 384: 193-205
- PubMed: 18824174 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.09.027
- Primary Citation Related Structures:
3E59, 3EAT - PubMed Abstract:
The pvcABCD operon of Pseudomonas aeruginosa encodes four proteins (PA2254, PA2255, PA2256, and PA2257) that form a cluster that is responsible for the synthesis of a cyclized isocyano derivative of tyrosine. These proteins, which were identified originally as being responsible for a step in the maturation of the chromophore of the peptide siderophore pyoverdine, have been identified recently as belonging to a family of proteins that produce small organic isonitriles. We report that strains harboring a disruption in the pvcA or pvcB genes are able to grow in iron-depleted conditions and to produce pyoverdine. Additionally, we have determined the three-dimensional crystal structures of PvcA and PvcB. The structure of PvcA demonstrates a novel enzyme architecture that is built upon a Rossmann fold. We have analyzed the sequence conservation of enzymes within this family and identified six conserved motifs. These regions of the protein cluster around a putative active site cavity. The structure of the PvcB protein confirms it is a member of the Fe2+/alpha-ketoglutarate-dependent oxygenase family of enzymes. The active site of PvcB is compared to the structures of other family members and suggests that a conformational change to order several loops will accompany the binding of ligands.
- Hauptman-Woodward Medical Research Institute, Department of Structural Biology, State University of New York at Buffalo, 700 Ellicott St, Buffalo, NY 14203-1102, USA.
Organizational Affiliation: