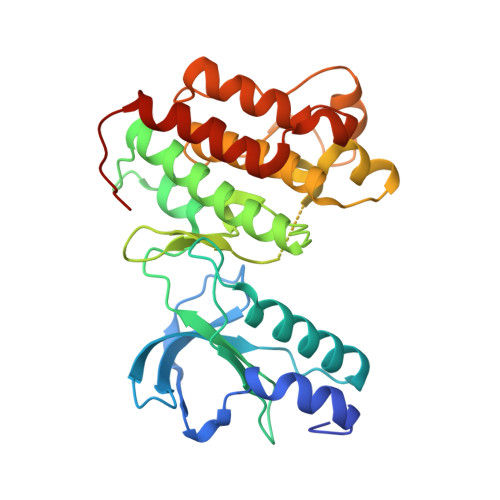
Design, synthesis, and biological evaluation of potent c-Met inhibitors.
D'Angelo, N.D., Bellon, S.F., Booker, S.K., Cheng, Y., Coxon, A., Dominguez, C., Fellows, I., Hoffman, D., Hungate, R., Kaplan-Lefko, P., Lee, M.R., Li, C., Liu, L., Rainbeau, E., Reider, P.J., Rex, K., Siegmund, A., Sun, Y., Tasker, A.S., Xi, N., Xu, S., Yang, Y., Zhang, Y., Burgess, T.L., Dussault, I., Kim, T.S.(2008) J Med Chem 51: 5766-5779
- PubMed: 18763753 Search on PubMed
- DOI: https://doi.org/10.1021/jm8006189
- Primary Citation Related Structures:
3EFJ, 3EFK - PubMed Abstract:
c-Met is a receptor tyrosine kinase that plays a key role in several cellular processes but has also been found to be overexpressed and mutated in different human cancers. Consequently, targeting this enzyme has become an area of intense research in drug discovery. Our studies began with the design and synthesis of novel pyrimidone 7, which was found to be a potent c-Met inhibitor. Subsequent SAR studies identified 22 as a more potent analog, whereas an X-ray crystal structure of 7 bound to c-Met revealed an unexpected binding conformation. This latter finding led to the development of a new series that featured compounds that were more potent both in vitro and in vivo than 22 and also exhibited different binding conformations to c-Met. Novel c-Met inhibitors have been designed, developed, and found to be potent in vitro and in vivo.
- Department of Medicinal Chemistry, Amgen Inc., One Amgen Center Drive, Thousand Oaks, California 91320-1799, USA. dangelo@amgen.com
Organizational Affiliation: