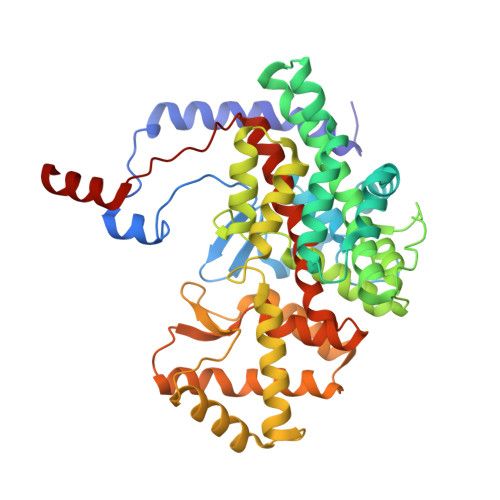
Structure of pig heart citrate synthase at 1.78 A resolution.
Larson, S.B., Day, J.S., Nguyen, C., Cudney, R., McPherson, A.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 430-434
- PubMed: 19407370 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109008343
- Primary Citation Related Structures:
3ENJ - PubMed Abstract:
Pig heart citrate synthase was crystallized from a small-molecule cocktail containing cystamine dihydrochloride, aspartame and benzamidine hydrochloride. The structure was refined to an R factor of 0.179 (R(free) = 0.222) using synchrotron data to a resolution of 1.78 A. The model includes the full-length protein, a chloride ion, a sulfate ion, 305 water molecules and an unexpected moiety attached through a disulfide linkage to Cys184, which was modeled as a half-cystamine molecule generated by disulfide exchange with the cystamine in the small-molecule cocktail.
- Department of Molecular Biology and Biochemistry, The University of California, Irvine, 92697-3900, USA.
Organizational Affiliation: