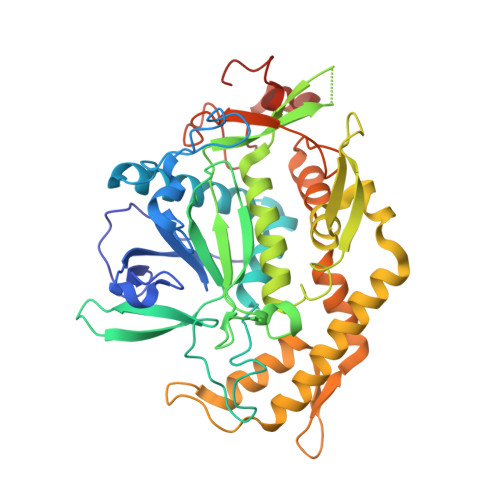
Mode of VAMP substrate recognition and inhibition of Clostridium botulinum neurotoxin F.
Agarwal, R., Schmidt, J.J., Stafford, R.G., Swaminathan, S.(2009) Nat Struct Mol Biol 16: 789-794
- PubMed: 19543288 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1626
- Primary Citation Related Structures:
3FIE, 3FII - PubMed Abstract:
Clostridium botulinum neurotoxins (BoNTs) cleave neuronal proteins responsible for neurotransmitter release, causing the neuroparalytic disease botulism. BoNT serotypes B, D, F and G cleave and inactivate vesicle-associated membrane protein (VAMP), each at a unique peptide bond. The specificity of BoNTs depends on the mode of substrate recognition. We have investigated the mechanism of substrate recognition of BoNT F by determining the crystal structures of its complex with two substrate-based inhibitors, VAMP 22-58/Gln58D-cysteine and 27-58/Gln58D-cysteine. The inhibitors bind to BoNT F in the canonical direction (as seen for BoNTs A and E substrates) but are positioned specifically via three major exosites away from the active site. The cysteine sulfur of the inhibitors interacts with the zinc and exists as sulfinic acid in the inhibitor VAMP 27-58/Gln58D-cysteine. Arg133 and Arg171, which form part of two separate exosites, are crucial for substrate binding and catalysis.
- Biology Department, Brookhaven National Laboratory, Upton, New York, USA.
Organizational Affiliation: