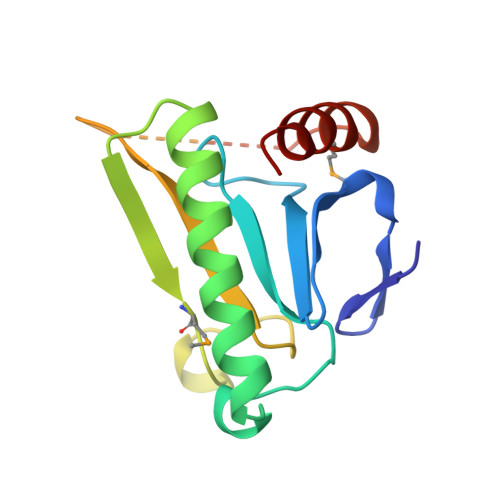
MAD analysis of FHIT, a putative human tumor suppressor from the HIT protein family.
Lima, C.D., D'Amico, K.L., Naday, I., Rosenbaum, G., Westbrook, E.M., Hendrickson, W.A.(1997) Structure 5: 763-774
- PubMed: 9261067 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(97)00231-1
- Primary Citation Related Structures:
1FIT, 2FIT, 3FIT - PubMed Abstract:
The fragile histidine triad (FHIT) protein is a member of the large and ubiquitous histidine triad (HIT) family of proteins. It is expressed from a gene located at a fragile site on human chromosome 3, which is commonly disrupted in association with certain cancers. On the basis of the genetic evidence, it has been postulated that the FHIT protein may function as a tumor suppressor, implying a role for the FHIT protein in carcinogenesis. The FHIT protein has dinucleoside polyphosphate hydrolase activity in vitro, thus suggesting that its role in vivo may involve the hydrolysis of a phosphoanhydride bond. The structural analysis of FHIT will identify critical residues involved in substrate binding and catalysis, and will provide insights into the in vivo function of HIT proteins. The three-dimensional crystal structures of free and nucleoside complexed FHIT have been determined from multiwavelength anomalous diffraction (MAD) data, and they represent some of the first successful structures to be measured with undulator radiation at the Advanced Photon Source. The structures of FHIT reveal that this protein exists as an intimate homodimer, which is based on a core structure observed previously in another human HIT homolog, protein kinase C interacting protein (PKCI), but has distinctive elaborations at both the N and C termini. Conserved residues within the HIT family, which are involved in the interactions of the proteins with nucleoside and phosphate groups, appear to be relevant for the catalytic activity of this protein. The structure of FHIT, a divergent HIT protein family member, in complex with a nucleotide analog suggests a metal-independent catalytic mechanism for the HIT family of proteins. A structural comparison of FHIT with PKCI and galactose-1-phosphate uridylyltransferase (GaIT) reveals additional implications for the structural and functional evolution of the ubiquitous HIT family of proteins.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: