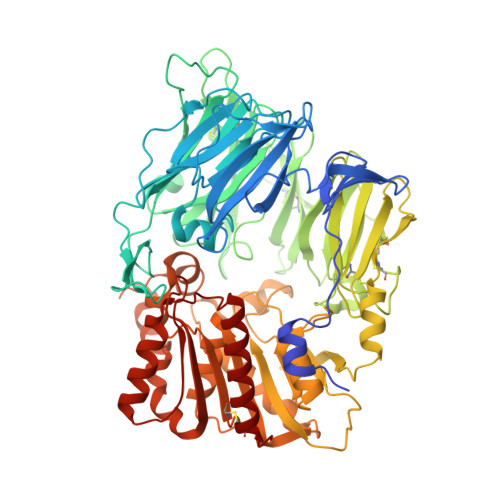
Design and Synthesis of Pyrimidinone and Pyrimidinedione Inhibitors of Dipeptidyl Peptidase IV.
Zhang, Z., Wallace, M.B., Feng, J., Stafford, J.A., Skene, R.J., Shi, L., Lee, B., Aertgeerts, K., Jennings, A., Xu, R., Kassel, D.B., Kaldor, S.W., Navre, M., Webb, D.R., Gwaltney, S.L.(2011) J Med Chem 54: 510-524
- PubMed: 21186796 Search on PubMed
- DOI: https://doi.org/10.1021/jm101016w
- Primary Citation Related Structures:
3G0B, 3G0C, 3G0D, 3G0G - PubMed Abstract:
The discovery of two classes of heterocyclic dipeptidyl peptidase IV (DPP-4) inhibitors, pyrimidinones and pyrimidinediones, is described. After a single oral dose, these potent, selective, and noncovalent inhibitors provide sustained reduction of plasma DPP-4 activity and lowering of blood glucose in animal models of diabetes. Compounds 13a, 27b, and 27j were selected for development.
- Takeda San Diego, Inc., 10410 Science Center Drive, San Diego, California 92121, United States.
Organizational Affiliation: