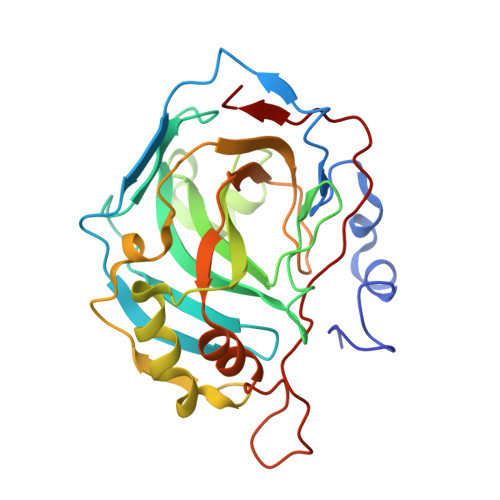
Apo-Human Carbonic Anhydrase II Revisited: Implications of the Loss of a Metal in Protein Structure, Stability, and Solvent Network .
Avvaru, B.S., Busby, S.A., Chalmers, M.J., Griffin, P.R., Venkatakrishnan, B., Agbandje-McKenna, M., Silverman, D.N., McKenna, R.(2009) Biochemistry 48: 7365-7372
- PubMed: 19583303 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi9007512
- Primary Citation Related Structures:
3GZ0 - PubMed Abstract:
Human carbonic anhydrase II (HCA II) is a monomeric zinc-containing metalloenzyme that catalyzes the hydration of CO(2) to form bicarbonate and a proton. The properties of the zinc have been extensively elucidated in catalysis but less well studied as a contributor to structure and stability. Apo-HCA II (without zinc) was prepared and compared to holo-HCA II: in crystallographic structural features, in backbone amide H/D exchange, and in thermal stability. The removal of zinc from the active site has no effect on either the topological fold of the enzyme or the ordered water network in the active site. However, the removal of the zinc alters the collective electrostatics of the apo-HCA II that result in the following differences from that of the holoenzyme: (1) the main thermal unfolding transition of the apo-HCA II is lowered by 8 degrees C, (2) the relative increase in thermal mobility of atoms of the apo-HCA II was not observed in the vicinity of the active site but manifested on the surface of the enzyme, and (3) the side chain of His 64, the proton shuttle residue that sits on the rim of the active site, is oriented outward and is associated with additional ordered "external" waters, as opposed to a near equal inward and outward orientation in the holo-HCA II.
- Department of Biochemistry and Molecular Biology, University of Florida, Gainesville, Florida 32610, USA.
Organizational Affiliation: