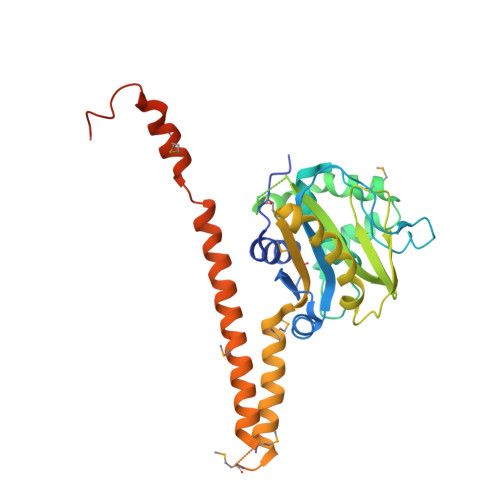
Structural characterization of human Uch37.
Burgie, S.E., Bingman, C.A., Soni, A.B., Phillips Jr., G.N.(2012) Proteins 80: 649-654
- PubMed: 21953935
- DOI: https://doi.org/10.1002/prot.23147
- Primary Citation of Related Structures:
3IHR - PubMed Abstract:
Uch37 is a de-ubiquitylating enzyme that is functionally linked with the 26S proteasome via Rpn13, and is essential for metazoan development. Here, we report the X-ray crystal structure of full-length human Uch37 at 2.95 Å resolution. Uch37's catalytic domain is similar to those of all UCH enzymes characterized to date. The C-terminal extension is elongated, predominantly helical and contains coiled coil interactions. Additionally, we provide an initial characterization of Uch37's oligomeric state and identify a systematic error in previous analyses of Uch37 activity. Taken together, these data provide a strong foundation for further analysis of Uch37's several functions.
- Department of Biochemistry, Center for Eukaryotic Structural Genomics, University of Wisconsin-Madison, Madison Wisconsin 53706-1544.
Organizational Affiliation: