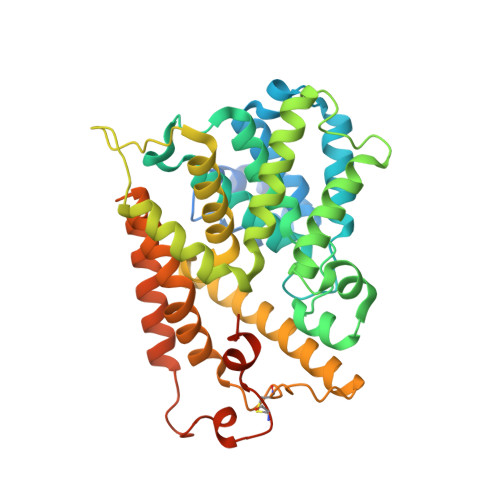
Crystal structure of human PDE4b with 5-[3-[(1S,2S,4R)-Bicyclo[2.2.1]hept-2-yloxy]-4-methoxyp henyl]tetrahydro-2(1H)-pyrimidinone reveals ordering of the C-terminal helix residues 502-509.
Cheng, R.K.Y., Crawley, L., Barker, J., Whittaker, M.To be published.