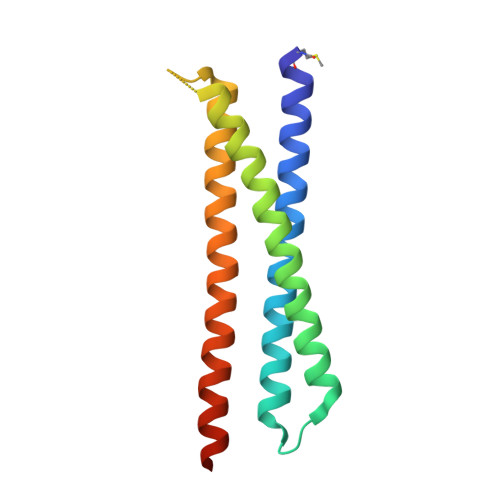
Computational Design of Epitope-Scaffolds Allows Induction of Antibodies Specific for a Poorly Immunogenic HIV Vaccine Epitope.
Correia, B.E., Ban, Y.E., Holmes, M.A., Xu, H., Ellingson, K., Kraft, Z., Carrico, C., Boni, E., Sather, D.N., Zenobia, C., Burke, K.Y., Bradley-Hewitt, T., Bruhn-Johannsen, J.F., Kalyuzhniy, O., Baker, D., Strong, R.K., Stamatatos, L., Schief, W.R.(2010) Structure 18: 1116-1126
- PubMed: 20826338 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2010.06.010
- Primary Citation Related Structures:
3LEF, 3LF6, 3LF9, 3LG7, 3LH2, 3LHP - PubMed Abstract:
Broadly cross-reactive monoclonal antibodies define epitopes for vaccine development against HIV and other highly mutable viruses. Crystal structures are available for several such antibody-epitope complexes, but methods are needed to translate that structural information into immunogens that re-elicit similar antibodies. We describe a general computational method to design epitope-scaffolds in which contiguous structural epitopes are transplanted to scaffold proteins for conformational stabilization and immune presentation. Epitope-scaffolds designed for the poorly immunogenic but conserved HIV epitope 4E10 exhibited high epitope structural mimicry, bound with higher affinities to monoclonal antibody (mAb) 4E10 than the cognate peptide, and inhibited HIV neutralization by HIV+ sera. Rabbit immunization with an epitope-scaffold induced antibodies with structural specificity highly similar to mAb 4E10, an important advance toward elicitation of neutralizing activity. The results demonstrate that computationally designed epitope-scaffolds are valuable as structure-specific serological reagents and as immunogens to elicit antibodies with predetermined structural specificity.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: