Structure-Function Analysis of the Short Splicing Variant Carboxypeptidase Encoded by Drosophila melanogaster silver.
Tanco, S., Arolas, J.L., Guevara, T., Lorenzo, J., Aviles, F.X., Gomis-Ruth, F.X.(2010) J Mol Biology 401: 465-477
- PubMed: 20600119 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.06.035
- Primary Citation Related Structures:
3MN8 - PubMed Abstract:
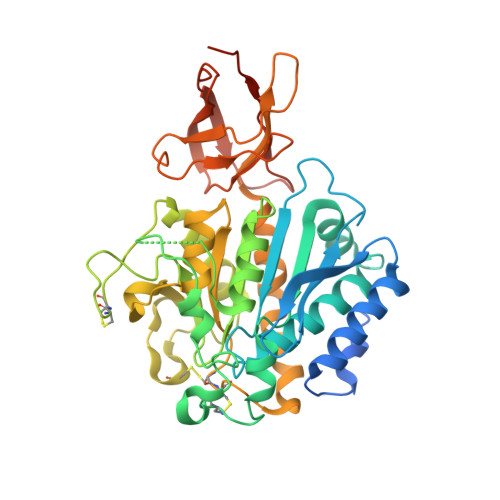
Drosophila melanogaster silver gene is the ortholog of the coding gene of mammalian carboxypeptidase D (CPD). The silver gene gives rise to eight different splicing variants of differing length that can contain up to three homologous repeats. Among the protein variants encoded, the short form 1B alias DmCPD1Bs (D. melanogaster CPD variant 1B short) is necessary and sufficient for viability of the fruit fly. It has one single repeat, it is active against standard peptide substrates, and it is localized to the secretory pathway. In this work, the enzyme was found as a monomer in solution and as a homodimer in the crystal structure, which features a protomer with an N-terminal 311-residue catalytic domain of alpha/beta-hydrolase fold and a C-terminal 84-residue all-beta transthyretin-like domain. Overall, DmCPD1Bs conforms to the structure of N/E-type funnelins/M14B metallopeptidases, but it has two unique structural elements potentially involved in regulation of its activity: (i) two contiguous surface cysteines that may become palmitoylated and target the enzyme to membranes, thus providing control through localization, and (ii) a surface hot spot targetable by peptidases that would provide a regulatory mechanism through proteolytic inactivation. Given that the fruit fly possesses orthologs of only two out of the five proteolytically competent N/E-type funnelins found in higher vertebrates, DmCPD1Bs may represent a functional analog of at least one of the missing mammalian CPs.
- Departament de Bioquímica i Biologia Molecular, Institut de Biotecnologia i de Biomedicina, Universitat Autònoma de Barcelona, E-08193 Bellaterra, Spain.
Organizational Affiliation: