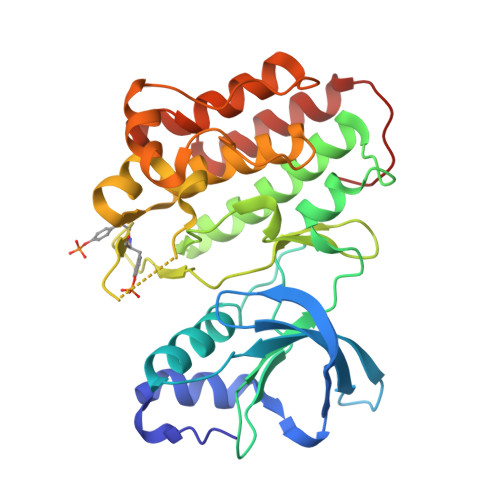
Structural basis for selective small molecule kinase inhibition of activated c-Met.
Rickert, K.W., Patel, S.B., Allison, T.J., Byrne, N.J., Darke, P.L., Ford, R.E., Guerin, D.J., Hall, D.L., Kornienko, M., Lu, J., Munshi, S.K., Reid, J.C., Shipman, J.M., Stanton, E.F., Wilson, K.J., Young, J.R., Soisson, S.M., Lumb, K.J.(2011) J Biological Chem 286: 11218-11225
- PubMed: 21247903 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.204404
- Primary Citation Related Structures:
3Q6U, 3Q6W, 3R7O - PubMed Abstract:
The receptor tyrosine kinase c-Met is implicated in oncogenesis and is the target for several small molecule and biologic agents in clinical trials for the treatment of cancer. Binding of the hepatocyte growth factor to the cell surface receptor of c-Met induces activation via autophosphorylation of the kinase domain. Here we describe the structural basis of c-Met activation upon autophosphorylation and the selective small molecule inhibiton of autophosphorylated c-Met. MK-2461 is a potent c-Met inhibitor that is selective for the phosphorylated state of the enzyme. Compound 1 is an MK-2461 analog with a 20-fold enthalpy-driven preference for the autophosphorylated over unphosphorylated c-Met kinase domain. The crystal structure of the unbound kinase domain phosphorylated at Tyr-1234 and Tyr-1235 shows that activation loop phosphorylation leads to the ejection and disorder of the activation loop and rearrangement of helix αC and the G loop to generate a viable active site. Helix αC adopts a orientation different from that seen in activation loop mutants. The crystal structure of the complex formed by the autophosphorylated c-Met kinase domain and compound 1 reveals a significant induced fit conformational change of the G loop and ordering of the activation loop, explaining the selectivity of compound 1 for the autophosphorylated state. The results highlight the role of structural plasticity within the kinase domain in imparting the specificity of ligand binding and provide the framework for structure-guided design of activated c-Met inhibitors.
- Global Structural Biology, Merck Research Laboratories, West Point, Pennsylvania 19486, USA.
Organizational Affiliation: