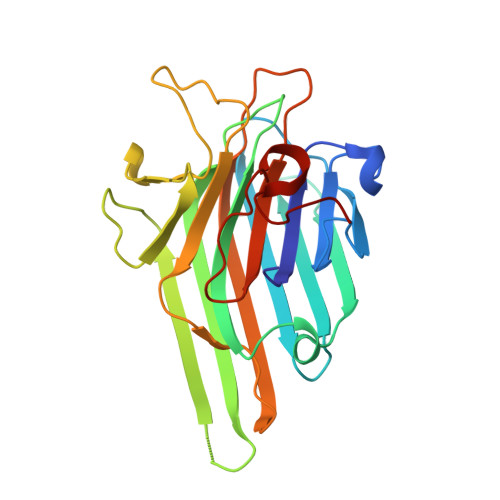
Synthesis and Biophysical Study of Disassembling Nano hybrid Bioconjugates with a Cubic Octasilsesquioxane Core
Trastoy, B., Bonsor, D.A., Perez-Ojeda, M.E., Sundberg, E.J., Jimeno, M.L., Garcia-Fernandez, J.M., Chiara, J.L.(2012) Adv Funct Mater 22: 3191-3201