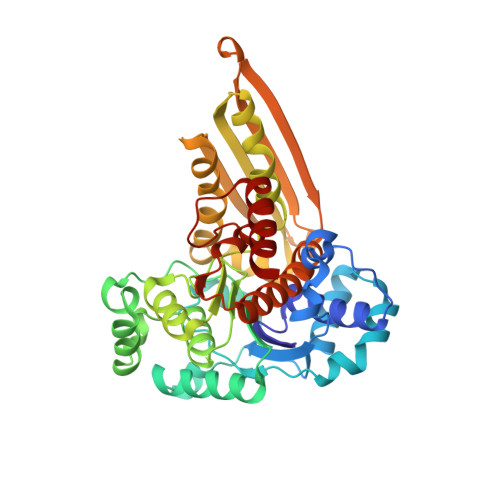
Crystal structure of a trapped catalytic intermediate suggests that forced atomic proximity drives the catalysis of mIPS.
Neelon, K., Roberts, M.F., Stec, B.(2011) Biophys J 101: 2816-2824
- PubMed: 22261071 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2011.10.038
- Primary Citation Related Structures:
3QVS, 3QVT, 3QVW, 3QVX, 3QW2 - PubMed Abstract:
1-L-myo-inositol-phosphate synthase (mIPS) catalyzes the first step of the unique, de novo pathway of inositol biosynthesis. However, details about the complex mIPS catalytic mechanism, which requires oxidation, enolization, intramolecular aldol cyclization, and reduction, are not fully known. To gain further insight into this mechanism, we determined the crystal structure of the wild-type mIPS from Archaeoglobus fulgidus at 1.7 Å, as well as the crystal structures of three active-site mutants. Additionally, we obtained the structure of mIPS with a trapped 5-keto-glucose-6-phosphate intermediate at 2 Å resolution by a novel (to our knowledge) process of activating the crystal at high temperature. A comparison of all of the crystal structures of mIPS described in this work suggests a novel type of catalytic mechanism that relies on the forced atomic proximity of functional groups. The lysine cluster is contained in a small volume in the active site, where random motions of these side chains are responsible for the progress of the complex multistep reaction as well as for the low rate of catalysis. The mechanism requires that functional groups of Lys-274, Lys-278, Lys-306, and Lys-367 assume differential roles in the protonation/deprotonation steps that must occur during the mIPS reaction. This mechanism is supported by the complete loss of activity of the enzyme caused by the Leu-257 mutation to Ala that releases the lysine containment.
- Department of Chemistry, Boston College, Chestnut Hill, Massachusetts, USA.
Organizational Affiliation: