Efficient and versatile one-step affinity purification of in vivo biotinylated proteins: Expression, characterization and structure analysis of recombinant human glutamate carboxypeptidase II.
Tykvart, J., Sacha, P., Barinka, C., Knedlik, T., Starkova, J., Lubkowski, J., Konvalinka, J.(2012) Protein Expr Purif 82: 106-115
- PubMed: 22178733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.pep.2011.11.016
- Primary Citation Related Structures:
3RBU - PubMed Abstract:
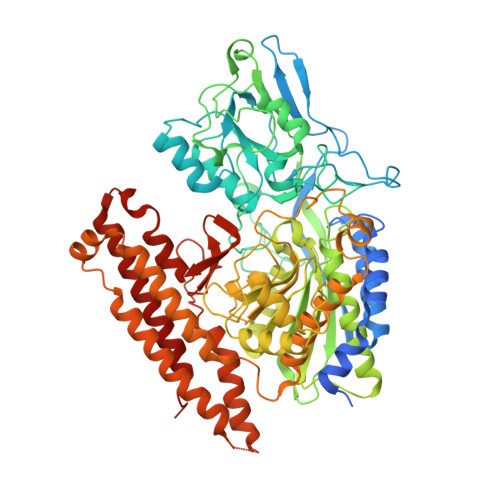
Affinity purification is a useful approach for purification of recombinant proteins. Eukaryotic expression systems have become more frequently used at the expense of prokaryotic systems since they afford recombinant eukaryotic proteins with post-translational modifications similar or identical to the native ones. Here, we present a one-step affinity purification set-up suitable for the purification of secreted proteins. The set-up is based on the interaction between biotin and mutated streptavidin. Drosophila Schneider 2 cells are chosen as the expression host, and a biotin acceptor peptide is used as an affinity tag. This tag is biotinylated by Escherichia coli biotin-protein ligase in vivo. We determined that localization of the ligase within the ER led to the most effective in vivo biotinylation of the secreted proteins. We optimized a protocol for large-scale expression and purification of AviTEV-tagged recombinant human glutamate carboxypeptidase II (Avi-GCPII) with milligram yields per liter of culture. We also determined the 3D structure of Avi-GCPII by X-ray crystallography and compared the enzymatic characteristics of the protein to those of its non-tagged variant. These experiments confirmed that AviTEV tag does not affect the biophysical properties of its fused partner. Purification approach, developed here, provides not only a sufficient amount of highly homogenous protein but also specifically and effectively biotinylates a target protein and thus enables its subsequent visualization or immobilization.
- Gilead Sciences and IOCB Research Centre, Institute of Organic Chemistry and Biochemistry, Academy of Sciences of the Czech Republic, Flemingovo n. 2, Prague 6, Czech Republic.
Organizational Affiliation: