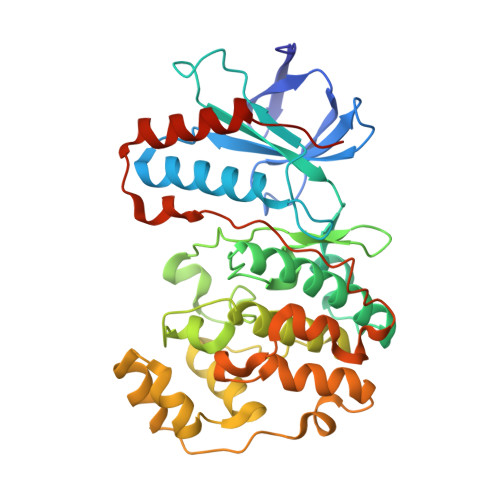
Indolin-2-one p38(alpha) inhibitors I: design, profiling and crystallographic binding mode.
Eastwood, P., Gonzalez, J., Gomez, E., Vidal, B., Caturla, F., Roca, R., Balague, C., Orellana, A., Dominguez, M.(2011) Bioorg Med Chem Lett 21: 4130-4133
- PubMed: 21696951 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.05.114
- Primary Citation Related Structures:
3RIN - PubMed Abstract:
The use of structure-based design and molecular modeling led to the discovery of indolin-2-one derivatives as potent and selective p38α inhibitors. The predicted binding mode was confirmed by X-ray crystallography.
- Almirall Research Center, Almirall S.A., Ctra. Laureà Miró 408, E-08980 St. Feliu de Llobregat, Barcelona, Spain. paul.eastwood@almirall.com
Organizational Affiliation: