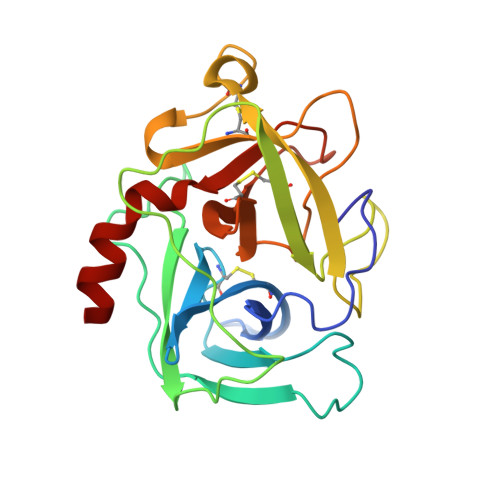
The structure of rat mast cell protease II at 1.9-A resolution.
Remington, S.J., Woodbury, R.G., Reynolds, R.A., Matthews, B.W., Neurath, H.(1988) Biochemistry 27: 8097-8105
- PubMed: 3233198 Search on PubMed
- DOI: https://doi.org/10.1021/bi00421a019
- Primary Citation Related Structures:
3RP2 - PubMed Abstract:
The structure of rat mast cell protease II (RMCP II), a serine protease with chymotrypsin-like primary specificity, has been determined to a nominal resolution of 1.9 A by single isomorphous replacement, molecular replacement, and restrained crystallographic refinement to a final R-factor of 0.191. There are two independent molecules of RMCP II in the asymmetric unit of the crystal. The rms deviation from ideal bond lengths is 0.016 A and from ideal bond angles is 2.7 degrees. The overall structure of RMCP II is extremely similar to that of chymotrypsin, but the largest differences between the two structures are clustered around the active-site region in a manner which suggests that the unusual substrate specificity of RMCP II is due to these changes. Unlike chymotrypsin, RMCP II has a deep cleft around the active site. An insertion of three residues between residues 35 and 41 of chymotrypsin, combined with concerted changes in sequence and a deletion near residue 61, allows residues 35-41 of RMCP II to adopt a conformation not seen in any other serine protease. Additionally, the loss of the disulfide bridge between residues 191 and 220 of chymotrypsin leads to the formation of an additional substrate binding pocket that we propose to interact with the P3 side chain of bound substrate. RMCP II is a member of a homologous subclass of serine proteases that are expressed by mast cells, neutrophils, lymphocytes, and cytotoxic T-cells. Thus, the structure of RMCP II forms a basis for an explanation of the unusual properties of other members of this class.
- Institute of Molecular Biology, University of Oregon, Eugene 97403.
Organizational Affiliation: