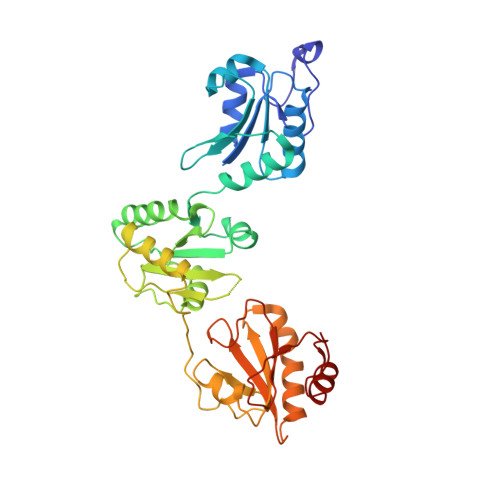
Human protein-disulfide isomerase is a redox-regulated chaperone activated by oxidation of domain a'
Wang, C., Yu, J., Huo, L., Wang, L., Feng, W., Wang, C.-C.(2012) J Biological Chem 287: 1139-1149
- PubMed: 22090031 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.303149
- Primary Citation Related Structures:
3UEM - PubMed Abstract:
Protein-disulfide isomerase (PDI), with domains arranged as abb'xa'c, is a key enzyme and chaperone localized in the endoplasmic reticulum (ER) catalyzing oxidative folding and preventing misfolding/aggregation of proteins. It has been controversial whether the chaperone activity of PDI is redox-regulated, and the molecular basis is unclear. Here, we show that both the chaperone activity and the overall conformation of human PDI are redox-regulated. We further demonstrate that the conformational changes are triggered by the active site of domain a', and the minimum redox-regulated cassette is located in b'xa'. The structure of the reduced bb'xa' reveals for the first time that domain a' packs tightly with both domain b' and linker x to form one compact structural module. Oxidation of domain a' releases the compact conformation and exposes the shielded hydrophobic areas to facilitate its high chaperone activity. Thus, the study unequivocally provides mechanistic insights into the redox-regulated chaperone activity of human PDI.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: