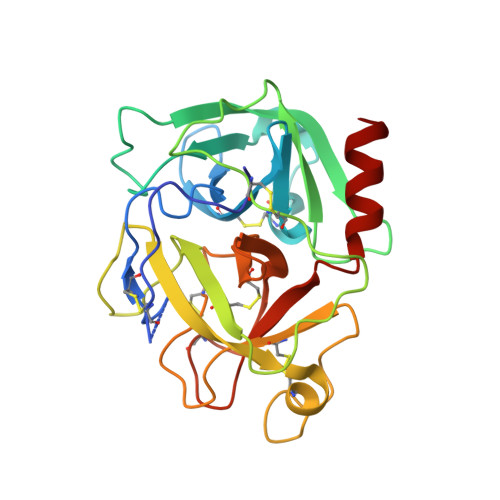
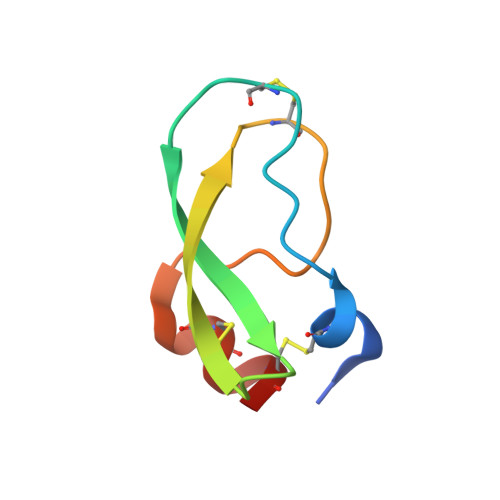
The structure of human microplasmin in complex with textilinin-1, an aprotinin-like inhibitor from the Australian brown snake.
Millers, E.K., Johnson, L.A., Birrell, G.W., Masci, P.P., Lavin, M.F., de Jersey, J., Guddat, L.W.(2013) PLoS One 8: e54104-e54104
- PubMed: 23335990 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0054104
- Primary Citation Related Structures:
3UIR - PubMed Abstract:
Textilinin-1 is a Kunitz-type serine protease inhibitor from Australian brown snake venom. Its ability to potently and specifically inhibit human plasmin (K(i) = 0.44 nM) makes it a potential therapeutic drug as a systemic anti-bleeding agent. The crystal structures of the human microplasmin-textilinin-1 and the trypsin-textilinin-1 complexes have been determined to 2.78 Å and 1.64 Å resolution respectively, and show that textilinin-1 binds to trypsin in a canonical mode but to microplasmin in an atypical mode with the catalytic histidine of microplasmin rotated out of the active site. The space vacated by the histidine side-chain in this complex is partially occupied by a water molecule. In the structure of microplasminogen the χ(1) dihedral angle of the side-chain of the catalytic histidine is rotated by 67° from its "active" position in the catalytic triad, as exemplified by its location when microplasmin is bound to streptokinase. However, when textilinin-1 binds to microplasmin the χ(1) dihedral angle of this amino acid residue changes by -157° (i.e. in the opposite rotation direction compared to microplasminogen). The unusual mode of interaction between textilinin-1 and plasmin explains textilinin-1's selectivity for human plasmin over plasma kallikrein. This difference can be exploited in future drug design efforts.
- School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, Queensland, Australia.
Organizational Affiliation: