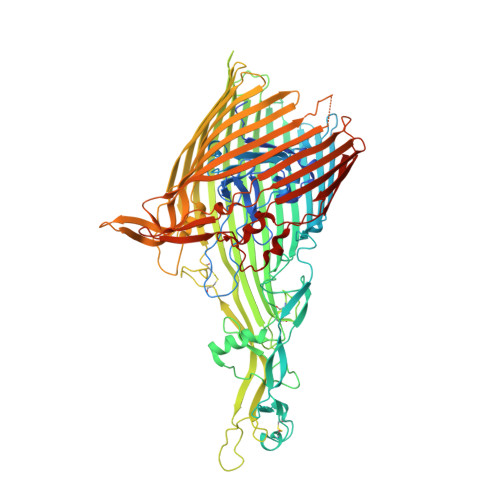
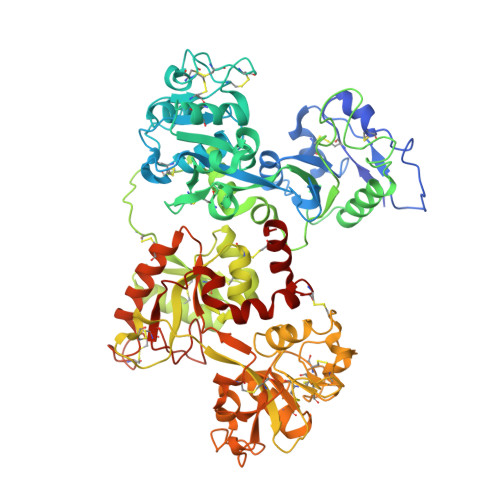
Structural basis for iron piracy by pathogenic Neisseria.
Noinaj, N., Easley, N.C., Oke, M., Mizuno, N., Gumbart, J., Boura, E., Steere, A.N., Zak, O., Aisen, P., Tajkhorshid, E., Evans, R.W., Gorringe, A.R., Mason, A.B., Steven, A.C., Buchanan, S.K.(2012) Nature 483: 53-58
- PubMed: 22327295 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature10823
- Primary Citation Related Structures:
3SKP, 3V83, 3V89, 3V8U, 3V8X - PubMed Abstract:
Neisseria are obligate human pathogens causing bacterial meningitis, septicaemia and gonorrhoea. Neisseria require iron for survival and can extract it directly from human transferrin for transport across the outer membrane. The transport system consists of TbpA, an integral outer membrane protein, and TbpB, a co-receptor attached to the cell surface; both proteins are potentially important vaccine and therapeutic targets. Two key questions driving Neisseria research are how human transferrin is specifically targeted, and how the bacteria liberate iron from transferrin at neutral pH. To address these questions, we solved crystal structures of the TbpA-transferrin complex and of the corresponding co-receptor TbpB. We characterized the TbpB-transferrin complex by small-angle X-ray scattering and the TbpA-TbpB-transferrin complex by electron microscopy. Our studies provide a rational basis for the specificity of TbpA for human transferrin, show how TbpA promotes iron release from transferrin, and elucidate how TbpB facilitates this process.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, US National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: