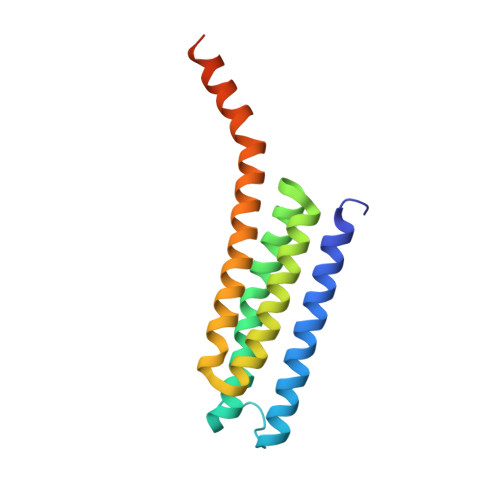
Seleno-detergent MAD phasing of leukotriene C4 synthase in complex with dodecyl-beta-D-selenomaltoside
Saino, H., Ago, H., Ukita, Y., Miyano, M.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1666-1673
- PubMed: 22139193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111042345
- Primary Citation Related Structures:
3B29 - PubMed Abstract:
Dodecyl-β-D-selenomaltoside (SeDDM) is a seleno-detergent with a β-glycosidic seleno-ether in place of the ether moiety in dodecyl-β-D-maltoside. Seleno-detergents are candidates for heavy-atom agents in experimental phasing of membrane proteins in protein crystallography. Crystals of a nuclear membrane-embedded enzyme, leukotriene C(4) synthase (LTC(4)S), in complex with SeDDM were prepared and a multiwavelength anomalous diffraction (MAD) experiment was performed. The SeDDM in the LTC(4)S crystal exhibited sufficient anomalous diffraction for determination of the structure using MAD phasing.
- Structural Biophysics Laboratory, RIKEN SPring-8 Center, Harima Institute, Sayo, Hyogo, Japan.
Organizational Affiliation: