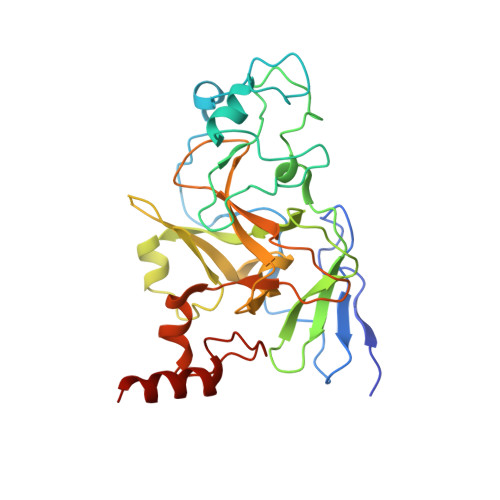
Discovery of a 2,4-diamino-7-aminoalkoxyquinazoline as a potent and selective inhibitor of histone lysine methyltransferase G9a.
Liu, F., Chen, X., Allali-Hassani, A., Quinn, A.M., Wasney, G.A., Dong, A., Barsyte, D., Kozieradzki, I., Senisterra, G., Chau, I., Siarheyeva, A., Kireev, D.B., Jadhav, A., Herold, J.M., Frye, S.V., Arrowsmith, C.H., Brown, P.J., Simeonov, A., Vedadi, M., Jin, J.(2009) J Med Chem 52: 7950-7953
- PubMed: 19891491 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm901543m
- Primary Citation Related Structures:
3K5K - PubMed Abstract:
SAR exploration of the 2,4-diamino-6,7-dimethoxyquinazoline template led to the discovery of 8 (UNC0224) as a potent and selective G9a inhibitor. A high resolution X-ray crystal structure of the G9a-8 complex, the first cocrystal structure of G9a with a small molecule inhibitor, was obtained. The cocrystal structure validated our binding hypothesis and will enable structure-based design of novel inhibitors. 8 is a useful tool for investigating the biology of G9a and its roles in chromatin remodeling.
- Eshelman School of Pharmacy, University of North Carolina, Chapel Hill, NC 27599, USA.
Organizational Affiliation: