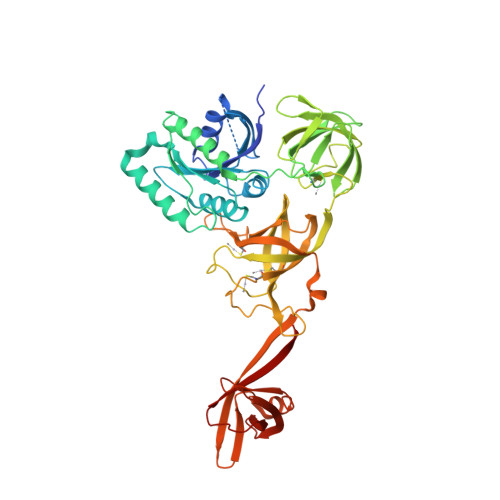Selenocysteine tRNA-Specific Elongation Factor Selb is a Structural Chimaera of Elongation and Initiation Factors.
Leibundgut, M., Frick, C., Thanbichler, M., Bock, A., Ban, N.(2005) EMBO J 24: 11
- PubMed: 15616587 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600505
- Primary Citation Related Structures:
4AC9, 4ACA, 4ACB - PubMed Abstract:
In all three kingdoms of life, SelB is a specialized translation elongation factor responsible for the cotranslational incorporation of selenocysteine into proteins by recoding of a UGA stop codon in the presence of a downstream mRNA hairpin loop. Here, we present the X-ray structures of SelB from the archaeon Methanococcus maripaludis in the apo-, GDP- and GppNHp-bound form and use mutational analysis to investigate the role of individual amino acids in its aminoacyl-binding pocket. All three SelB structures reveal an EF-Tu:GTP-like domain arrangement. Upon binding of the GTP analogue GppNHp, a conformational change of the Switch 2 region in the GTPase domain leads to the exposure of SelB residues involved in clamping the 5' phosphate of the tRNA. A conserved extended loop in domain III of SelB may be responsible for specific interactions with tRNA(Sec) and act as a ruler for measuring the extra long acceptor arm. Domain IV of SelB adopts a beta barrel fold and is flexibly tethered to domain III. The overall domain arrangement of SelB resembles a 'chalice' observed so far only for initiation factor IF2/eIF5B. In our model of SelB bound to the ribosome, domain IV points towards the 3' mRNA entrance cleft ready to interact with the downstream secondary structure element.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Zürich, Zürich, Switzerland.
Organizational Affiliation: