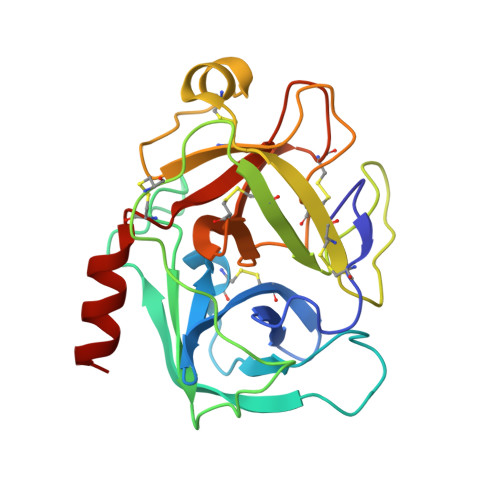
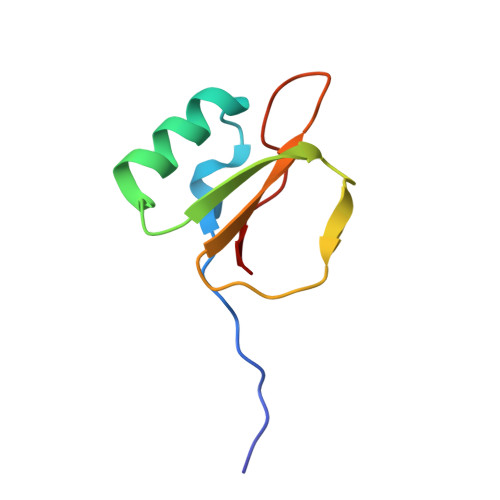
Thermodynamic signatures in macromolecular interactions involving conformational flexibility.
Menzel, A., Neumann, P., Schwieger, C., Stubbs, M.T.(2014) Biol Chem 395: 905-911
- PubMed: 25003391 Search on PubMed
- DOI: https://doi.org/10.1515/hsz-2014-0177
- Primary Citation Related Structures:
4B1T, 4B2A, 4B2B, 4B2C - PubMed Abstract:
The energetics of macromolecular interactions are complex, particularly where protein flexibility is involved. Exploiting serendipitous differences in the plasticity of a series of closely related trypsin variants, we analyzed the enthalpic and entropic contributions accompanying interaction with L45K-eglin C. Binding of the four variants show significant differences in released heat, although the affinities vary little, in accordance with the principle of enthalpy-entropy compensation. Binding of the most disordered variant is almost entirely enthalpically driven, with practically no entropy change. As structures of the complexes reveal negligible differences in protein-inhibitor contacts, we conclude that solvent effects contribute significantly to binding affinities.