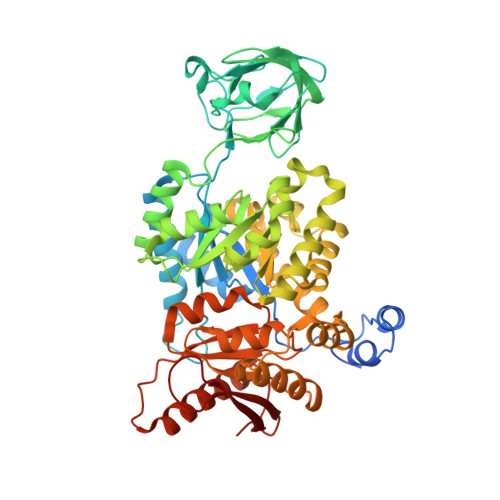
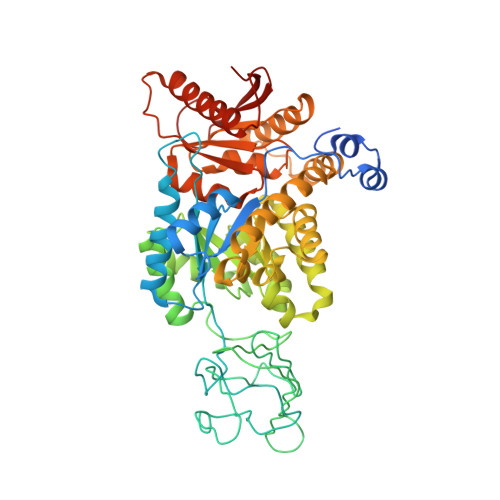
Serine is a natural ligand and allosteric activator of pyruvate kinase M2.
Chaneton, B., Hillmann, P., Zheng, L., Martin, A.C.L., Maddocks, O.D.K., Chokkathukalam, A., Coyle, J.E., Jankevics, A., Holding, F.P., Vousden, K.H., Frezza, C., O'Reilly, M., Gottlieb, E.(2012) Nature 491: 458-462
- PubMed: 23064226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature11540
- Primary Citation Related Structures:
4B2D - PubMed Abstract:
Cancer cells exhibit several unique metabolic phenotypes that are critical for cell growth and proliferation. Specifically, they overexpress the M2 isoform of the tightly regulated enzyme pyruvate kinase (PKM2), which controls glycolytic flux, and are highly dependent on de novo biosynthesis of serine and glycine. Here we describe a new rheostat-like mechanistic relationship between PKM2 activity and serine biosynthesis. We show that serine can bind to and activate human PKM2, and that PKM2 activity in cells is reduced in response to serine deprivation. This reduction in PKM2 activity shifts cells to a fuel-efficient mode in which more pyruvate is diverted to the mitochondria and more glucose-derived carbon is channelled into serine biosynthesis to support cell proliferation.
- Cancer Research UK, The Beatson Institute for Cancer Research, Switchback Road, Glasgow, G61 1BD, Scotland, United Kingdom.
Organizational Affiliation: