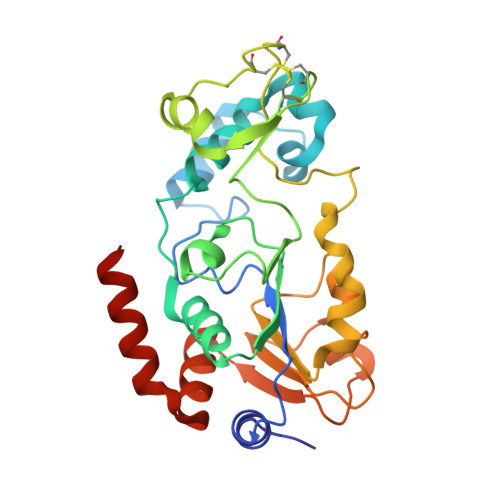
Structures of Human Sirtuin 3 Complexes with Adp-Ribose and with Carba-Nad+ and Srt1720: Binding Details and Inhibition Mechanism
Nguyen, G.T.T., Schaefer, S., Gertz, M., Weyand, M., Steegborn, C.(2013) Acta Crystallogr D Biol Crystallogr 69: 1423
- PubMed: 23897466 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913015448
- Primary Citation Related Structures:
4BN4, 4BN5 - PubMed Abstract:
Sirtuins are NAD(+)-dependent protein deacetylases that regulate metabolism and aging processes and are considered to be attractive therapeutic targets. Most available sirtuin modulators are little understood mechanistically, hindering their improvement. SRT1720 was initially described as an activator of human Sirt1, but it also potently inhibits human Sirt3. Here, the molecular mechanism of the inhibition of Sirt3 by SRT1720 is described. A crystal structure of Sirt3 in complex with SRT1720 and an NAD(+) analogue reveals that the compound partially occupies the acetyl-Lys binding site, thus explaining the reported competition with the peptide substrate. The compound packs against a hydrophobic protein patch and binds with its opposite surface to the NAD(+) nicotinamide, resulting in an exceptionally tight sandwich-like interaction. The observed arrangement rationalizes the uncompetitive inhibition with NAD(+), and binding measurements confirm that the nicotinamide moiety of NAD(+) supports inhibitor binding. Consistently, no inhibitor is bound in a second crystal structure of Sirt3 that was solved complexed with ADP-ribose and crystallized in the presence of SRT1720. These results reveal a novel sirtuin inhibitor binding site and mechanism, and provide a structural basis for compound improvement.
- Department of Biochemistry, University of Bayreuth, Universitätsstrasse 30, 95445 Bayreuth, Germany.
Organizational Affiliation: