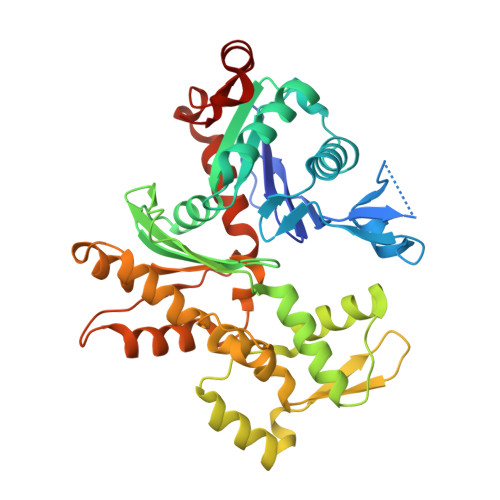
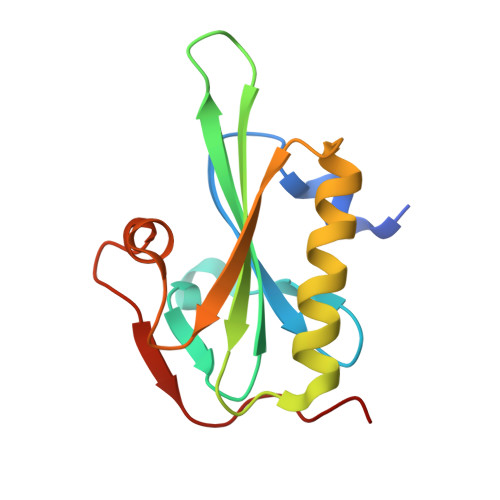
Structural Differences Explain Diverse Functions of Plasmodium Actins.
Vahokoski, J., Bhargav, S.P., Desfosses, A., Andreadaki, M., Kumpula, E., Martinez, S.M., Ignatev, A., Lepper, S., Frischknecht, F., Siden-Kiamos, I., Sachse, C., Kursula, I.(2014) PLoS Pathog 10: 4091
- PubMed: 24743229 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1004091
- Primary Citation Related Structures:
4CBU, 4CBW, 4CBX - PubMed Abstract:
Actins are highly conserved proteins and key players in central processes in all eukaryotic cells. The two actins of the malaria parasite are among the most divergent eukaryotic actins and also differ from each other more than isoforms in any other species. Microfilaments have not been directly observed in Plasmodium and are presumed to be short and highly dynamic. We show that actin I cannot complement actin II in male gametogenesis, suggesting critical structural differences. Cryo-EM reveals that Plasmodium actin I has a unique filament structure, whereas actin II filaments resemble canonical F-actin. Both Plasmodium actins hydrolyze ATP more efficiently than α-actin, and unlike any other actin, both parasite actins rapidly form short oligomers induced by ADP. Crystal structures of both isoforms pinpoint several structural changes in the monomers causing the unique polymerization properties. Inserting the canonical D-loop to Plasmodium actin I leads to the formation of long filaments in vitro. In vivo, this chimera restores gametogenesis in parasites lacking actin II, suggesting that stable filaments are required for exflagellation. Together, these data underline the divergence of eukaryotic actins and demonstrate how structural differences in the monomers translate into filaments with different properties, implying that even eukaryotic actins have faced different evolutionary pressures and followed different paths for developing their polymerization properties.
- Faculty of Biochemistry and Molecular Medicine, University of Oulu, Oulu, Finland.
Organizational Affiliation: