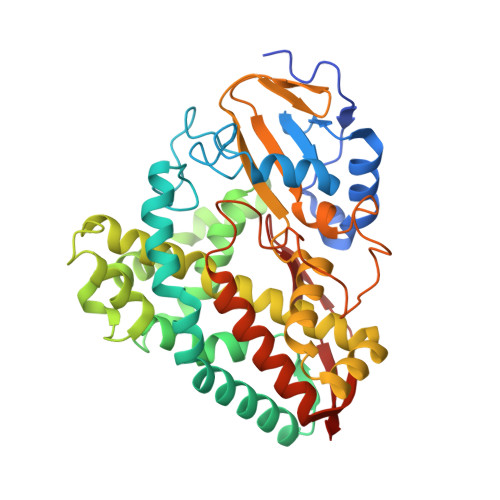
The crystal structures of 4-methoxybenzoate bound CYP199A2 and CYP199A4: structural changes on substrate binding and the identification of an anion binding site
Bell, S.G., Yang, W., Tan, A.B.H., Zhou, R., Johnson, E.O.D., Zhang, A., Zhou, W., Rao, Z., Wong, L.-L.(2012) Dalton Trans 41: 8703-8714
- PubMed: 22695988 Search on PubMed
- DOI: https://doi.org/10.1039/c2dt30783a
- Primary Citation Related Structures:
4DNJ, 4DNZ, 4DO1 - PubMed Abstract:
The crystal structures of the 4-methoxybenzoate bound forms of cytochrome P450 enzymes CYP199A2 and CYP199A4 from the Rhodopseudomonas palustris strains CGA009 and HaA2 have been solved. The structures of these two enzymes, which share 86% sequence identity, are very similar though some differences are found on the proximal surface. In these structures the enzymes have a closed conformation, in contrast to the substrate-free form of CYP199A2 where an obvious substrate access channel is observed. The switch from an open to a closed conformation arises from pronounced residue side-chain movements and alterations of ion pair and hydrogen bonding interactions at the entrance of the access channel. A chloride ion bound just inside the protein surface caps the entrance to the active site and protects the substrate and the heme from the external solvent. In both structures the substrate is held in place via hydrophobic and hydrogen bond interactions. The methoxy group is located over the heme iron, accounting for the high activity and selectivity of these enzymes for oxidative demethylation of the substrate. Mutagenesis studies on CYP199A4 highlight the involvement of hydrophobic (Phe185) and hydrophilic (Arg92, Ser95 and Arg243) amino acid residues in the binding of para-substituted benzoates by these enzymes.
- Department of Chemistry, University of Oxford, Inorganic Chemistry Laboratory, South Parks Road, Oxford, OX1 3QR, UK. stephen.bell@adelaide.edu.au
Organizational Affiliation: