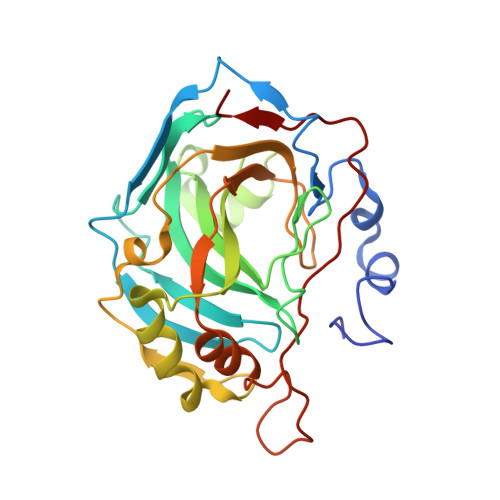
Hydroxamate represents a versatile zinc binding group for the development of new carbonic anhydrase inhibitors.
Di Fiore, A., Maresca, A., Supuran, C.T., De Simone, G.(2012) Chem Commun (Camb) 48: 8838-8840
- PubMed: 22836518 Search on PubMed
- DOI: https://doi.org/10.1039/c2cc34275h
- Primary Citation Related Structures:
4FL7 - PubMed Abstract:
Hydroxamates (R-CONHOH) have been scarcely investigated as carbonic anhydrase (CA, EC 4.2.1.1) inhibitors (CAIs). An inhibition/structural study of PhCONHOH is reported against all human isoforms. Comparing aliphatic (R = Me and CF(3)) and aromatic (R = Ph) hydroxamates as CAIs, we prove that CONHOH is a versatile zinc binding group. Depending on the nature of the R moiety, it can adopt different coordination modes to the catalytic ion within the CA active site.
- Istituto di Biostrutture e Bioimmagini - CNR, Napoli, Italy.
Organizational Affiliation: