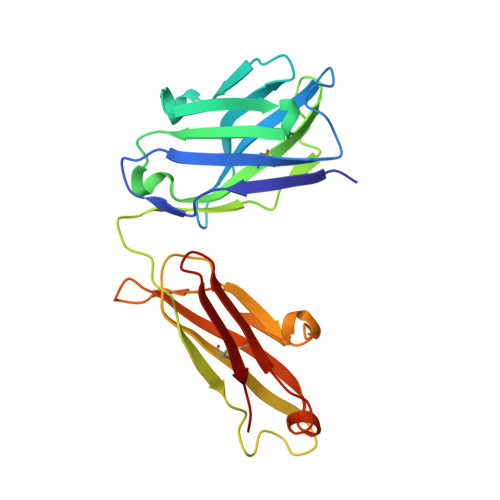
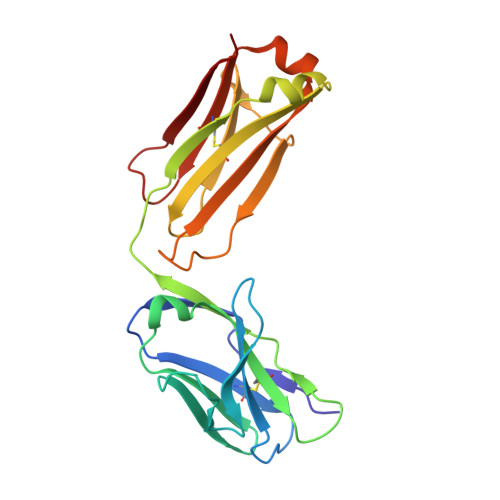
One target-two different binding modes: Structural insights into gevokizumab and canakinumab interactions to interleukin-1beta
Blech, M., Peter, D., Fischer, P., Bauer, M.M., Hafner, M., Zeeb, M., Nar, H.(2013) J Mol Biology 425: 94-111
- PubMed: 23041424 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.09.021
- Primary Citation Related Structures:
4G5Z, 4G6J, 4G6K, 4G6M - PubMed Abstract:
Interleukin-1β (IL-1β) is a key orchestrator in inflammatory and several immune responses. IL-1β exerts its effects through interleukin-1 receptor type I (IL-1RI) and interleukin-1 receptor accessory protein (IL-1RAcP), which together form a heterotrimeric signaling-competent complex. Canakinumab and gevokizumab are highly specific IL-1β monoclonal antibodies. Canakinumab is known to neutralize IL-1β by competing for binding to IL-1R and therefore blocking signaling by the antigen:antibody complex. Gevokizumab is claimed to be a regulatory therapeutic antibody that modulates IL-1β bioactivity by reducing the affinity for its IL-1RI:IL-1RAcP signaling complex. How IL-1β signaling is affected by both canakinumab and gevokizumab was not yet experimentally determined. We have analyzed the crystal structures of canakinumab and gevokizumab antibody binding fragment (Fab) as well as of their binary complexes with IL-1β. Furthermore, we characterized the epitopes on IL-1β employed by the antibodies by NMR epitope mapping studies. The direct comparison of NMR and X-ray data shows that the epitope defined by the crystal structure encompasses predominantly those residues whose NMR resonances are severely perturbed upon complex formation. The antigen:Fab co-structures confirm the previously identified key contact residues on IL-1β and provide insight into the mechanisms leading to their distinct modulation of IL-1β signaling. A significant steric overlap of the binding interfaces of IL-1R and canakinumab on IL-1β causes competitive inhibition of the association of IL-1β and its receptor. In contrast, gevokizumab occupies an allosteric site on IL-1β and complex formation results in a minor reduction of binding affinity to IL-1RI. This suggests two different mechanisms of IL-1β pathway attenuation.
- Department of Lead Identification and Optimization Support, Structural Research Group, Boehringer Ingelheim Pharma GmbH & Co. KG, Birkendorfer Str. 65, 88397 Biberach, Germany. michaela.blech@boehringer-ingelheim.com
Organizational Affiliation: