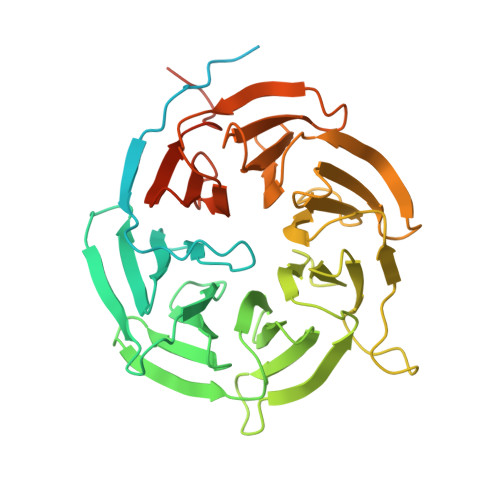
Structural analysis of human Cdc20 supports multisite degron recognition by APC/C.
Tian, W., Li, B., Warrington, R., Tomchick, D.R., Yu, H., Luo, X.(2012) Proc Natl Acad Sci U S A 109: 18419-18424
- PubMed: 23091007 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1213438109
- Primary Citation Related Structures:
4GGA, 4GGC, 4GGD - PubMed Abstract:
The anaphase-promoting complex/cyclosome (APC/C) promotes anaphase onset and mitotic exit through ubiquitinating securin and cyclin B1. The mitotic APC/C activator, the cell division cycle 20 (Cdc20) protein, directly interacts with APC/C degrons--the destruction (D) and KEN boxes. APC/C(Cdc20) is the target of the spindle checkpoint. Checkpoint inhibition of APC/C(Cdc20) requires the binding of a BubR1 KEN box to Cdc20. How APC/C recognizes substrates is not understood. We report the crystal structures of human Cdc20 alone or bound to a BubR1 KEN box. Cdc20 has a disordered N-terminal region and a C-terminal WD40 β propeller with a preformed KEN-box-binding site at its top face. We identify a second conserved surface at the side of the Cdc20 β propeller as a D-box-binding site. The D box of securin, but not its KEN box, is critical for securin ubiquitination by APC/C(Cdc20). Although both motifs contribute to securin ubiquitination by APC/C(Cdh1), securin mutants lacking either motif are efficiently ubiquitinated. Furthermore, D-box peptides diminish the ubiquitination of KEN-box substrates by APC/C(Cdh1), suggesting possible competition between the two motifs. Our results indicate the lack of strong positive cooperativity between the two degrons of securin. We propose that low-cooperativity, multisite target recognition enables APC/C to robustly ubiquitinate diverse substrates and helps to drive cell cycle oscillations.
- Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: