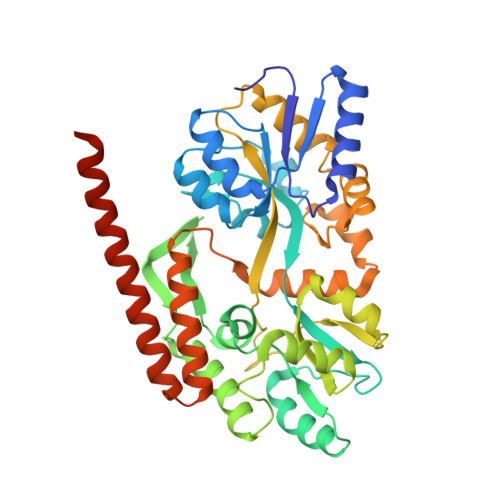
The survival motor neuron protein forms soluble glycine zipper oligomers.
Martin, R., Gupta, K., Ninan, N.S., Perry, K., Van Duyne, G.D.(2012) Structure 20: 1929-1939
- PubMed: 23022347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.08.024
- Primary Citation Related Structures:
4GLI - PubMed Abstract:
The survival motor neuron (SMN) protein forms the oligomeric core of a multiprotein complex that functions in spliceosomal snRNP biogenesis. Loss of function mutations in the SMN gene cause spinal muscular atrophy (SMA), a leading genetic cause of infant mortality. Nearly half of the known SMA patient missense mutations map to the SMN YG-box, a highly conserved oligomerization domain of unknown structure that contains a (YxxG)₃ motif. Here, we report that the SMN YG-box forms helical oligomers similar to the glycine zippers found in transmembrane channel proteins. A network of tyrosine-glycine packing between helices drives formation of soluble YG-box oligomers, providing a structural basis for understanding SMN oligomerization and for relating defects in oligomerization to the mutations found in SMA patients. These results have important implications for advancing our understanding of SMN function and glycine zipper-mediated helix-helix interactions.
- Graduate Group in Biochemistry and Molecular Biophysics, University of Pennsylvania, Philadelphia, PA 19104, USA.
Organizational Affiliation: