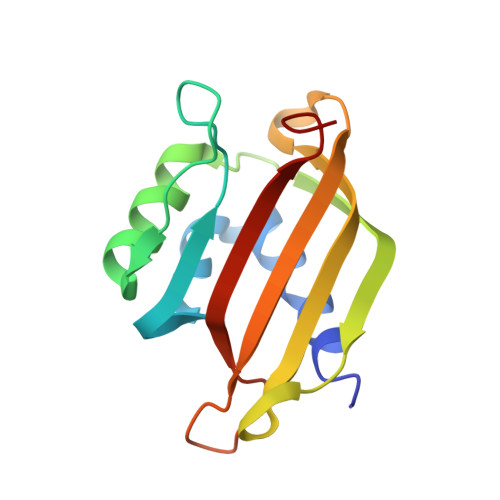
Crystal Structures of the Human G3BP1 NTF2-Like Domain Visualize FxFG Nup Repeat Specificity.
Vognsen, T., Moller, I.R., Kristensen, O.(2013) PLoS One 8: e80947-e80947
- PubMed: 24324649 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0080947
- Primary Citation Related Structures:
4FCJ, 4FCM, 4IIA - PubMed Abstract:
Ras GTPase Activating Protein SH3 Domain Binding Protein (G3BP) is a potential anti-cancer drug target implicated in several cellular functions. We have used protein crystallography to solve crystal structures of the human G3BP1 NTF2-like domain both alone and in complex with an FxFG Nup repeat peptide. Despite high structural similarity, the FxFG binding site is located between two alpha helices in the G3BP1 NTF2-like domain and not at the dimer interface as observed for nuclear transport factor 2. ITC studies showed specificity towards the FxFG motif but not FG and GLFG motifs. The unliganded form of the G3BP1 NTF2-like domain was solved in two crystal forms to resolutions of 1.6 and 3.3 Å in space groups P212121 and P6322 based on two different constructs, residues 1-139 and 11-139, respectively. Crystal packing of the N-terminal residues against a symmetry related molecule in the P212121 crystal form might indicate a novel ligand binding site that, however, remains to be validated. The crystal structures give insight into the nuclear transportation mechanisms of G3BP and provide a basis for future structure based drug design.
- Biostructural Research, Department of Drug Design and Pharmacology, Faculty of Health Sciences, University of Copenhagen, Copenhagen, Denmark.
Organizational Affiliation: