Discovery of potent and efficacious urea-containing nicotinamide phosphoribosyltransferase (NAMPT) inhibitors with reduced CYP2C9 inhibition properties.
Gunzner-Toste, J., Zhao, G., Bauer, P., Baumeister, T., Buckmelter, A.J., Caligiuri, M., Clodfelter, K.H., Fu, B., Han, B., Ho, Y.C., Kley, N., Liang, X., Liederer, B.M., Lin, J., Mukadam, S., O'Brien, T., Oh, A., Reynolds, D.J., Sharma, G., Skelton, N., Smith, C.C., Sodhi, J., Wang, W., Wang, Z., Xiao, Y., Yuen, P.W., Zak, M., Zhang, L., Zheng, X., Bair, K.W., Dragovich, P.S.(2013) Bioorg Med Chem Lett 23: 3531-3538
- PubMed: 23668988 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2013.04.040
- Primary Citation Related Structures:
4JNM - PubMed Abstract:
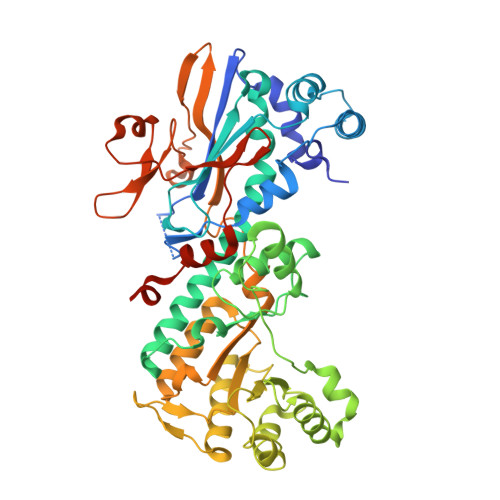
Potent, reversible inhibition of the cytochrome P450 CYP2C9 isoform was observed in a series of urea-containing nicotinamide phosphoribosyltransferase (NAMPT) inhibitors. This unwanted property was successfully removed from the described inhibitors through a combination of structure-based design and medicinal chemistry activities. An optimized compound which did not inhibit CYP2C9 exhibited potent anti-NAMPT activity (17; BC NAMPT IC50=3 nM; A2780 antiproliferative IC50=70 nM), good mouse PK properties, and was efficacious in an A2780 mouse xenograft model. The crystal structure of this compound in complex with the NAMPT protein is also described.
- Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: