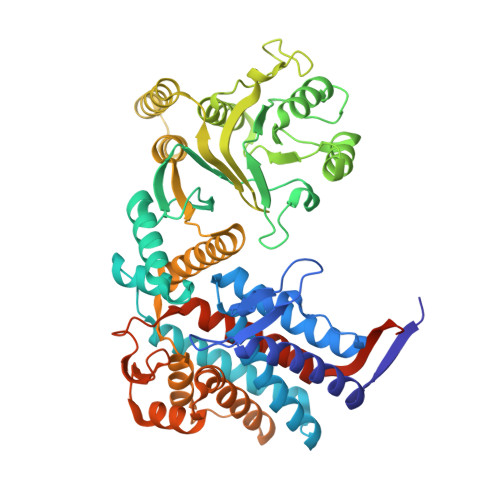
Crystal structure of a GroEL-ADP complex in the relaxed allosteric state at 2.7 A resolution.
Fei, X., Yang, D., Laronde-Leblanc, N., Lorimer, G.H.(2013) Proc Natl Acad Sci U S A 110: E2958-E2966
- PubMed: 23861496
- DOI: https://doi.org/10.1073/pnas.1311996110
- Primary Citation of Related Structures:
4KI8 - PubMed Abstract:
The chaperonin proteins GroEL and GroES are cellular nanomachines driven by the hydrolysis of ATP that facilitate the folding of structurally diverse substrate proteins. In response to ligand binding, the subunits of a ring cycle in a concerted manner through a series of allosteric states (T, R, and R″), enabling work to be performed on the substrate protein. Removing two salt bridges that ordinarily break during the allosteric transitions of the WT permitted the structure of GroEL-ADP in the R state to be solved to 2.7 Å resolution. Whereas the equatorial domain displays almost perfect sevenfold symmetry, the apical domains, to which substrate proteins bind, and to a lesser extent, the intermediate domains display a remarkable asymmetry. Freed of intersubunit contacts, the apical domain of each subunit adopts a different conformation, suggesting a flexibility that permits interaction with diverse substrate proteins. This result contrasts with a previous cryo-EM study of a related allosteric ATP-bound state at lower resolution. After artificially imposing sevenfold symmetry it was concluded that a GroEL ring in the R-ATP state existed in six homogeneous but slightly different states. By imposing sevenfold symmetry on each of the subunits of the crystal structure of GroEL-ADP, we showed that the synthetic rings of (X-ray) GroEL-ADP and (cryo-EM) GroEL-ATP are structurally closely related. A deterministic model, the click stop mechanism, that implied temporal transitions between these states was proposed. Here, however, these conformational states are shown to exist as a structurally heterogeneous ensemble within a single ring.
- Center for Biological Structure and Organization, University of Maryland, College Park, MD 20742, USA.
Organizational Affiliation: