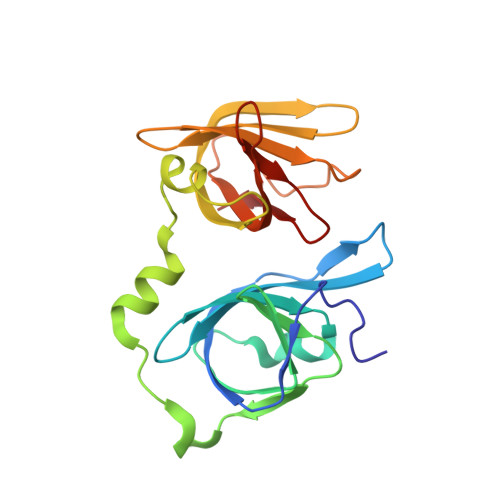
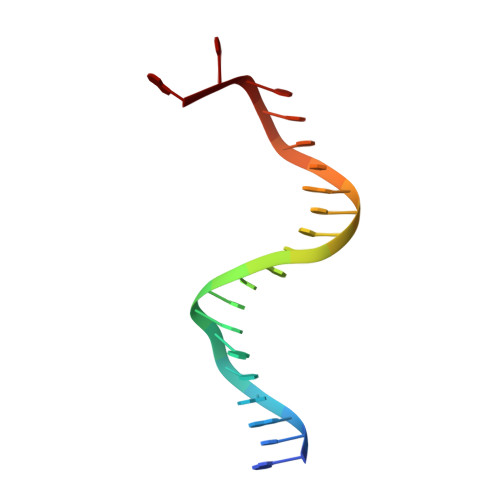
Molecular Mechanism for p202-Mediated Specific Inhibition of AIM2 Inflammasome Activation.
Yin, Q., Sester, D.P., Tian, Y., Hsiao, Y.S., Lu, A., Cridland, J.A., Sagulenko, V., Thygesen, S.J., Choubey, D., Hornung, V., Walz, T., Stacey, K.J., Wu, H.(2013) Cell Rep 4: 327-339
- PubMed: 23850291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2013.06.024
- Primary Citation Related Structures:
4L5Q, 4L5R, 4L5S, 4L5T - PubMed Abstract:
Mouse p202 containing two hemopoietic expression, interferon inducibility, nuclear localization (HIN) domains antagonizes AIM2 inflammasome signaling and potentially modifies lupus susceptibility. We found that only HIN1 of p202 binds double-stranded DNA (dsDNA), while HIN2 forms a homotetramer. Crystal structures of HIN1 revealed that dsDNA is bound on face opposite the site used in AIM2 and IFI16. The structure of HIN2 revealed a dimer of dimers, the face analogous to the HIN1 dsDNA binding site being a dimerization interface. Electron microscopy imaging showed that HIN1 is flexibly linked to HIN2 in p202, and tetramerization provided enhanced avidity for dsDNA. Surprisingly, HIN2 of p202 interacts with the AIM HIN domain. We propose that this results in a spatial separation of the AIM2 pyrin domains, and indeed p202 prevented the dsDNA-dependent clustering of apoptosis-associated speck-like protein containing caspase recruitment domain (ASC) and AIM2 inflammasome activation. We hypothesize that while p202 was evolutionarily selected to limit AIM2-mediated inflammation in some mouse strains, the same mechanism contributes to increased interferon production and lupus susceptibility.
- Program in Cellular and Molecular Medicine, Boston Children's Hospital, and Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: