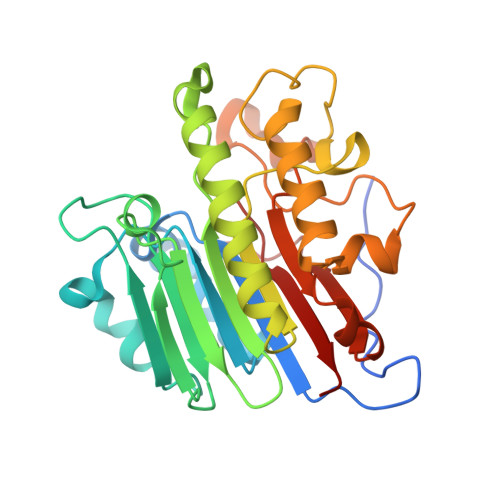
Structure of human apurinic/apyrimidinic endonuclease 1 with the essential Mg(2+) cofactor.
Manvilla, B.A., Pozharski, E., Toth, E.A., Drohat, A.C.(2013) Acta Crystallogr D Biol Crystallogr 69: 2555-2562
- PubMed: 24311596 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913027042
- Primary Citation Related Structures:
4LND - PubMed Abstract:
Apurinic/apyrimidinic endonuclease 1 (APE1) mediates the repair of abasic sites and other DNA lesions and is essential for base-excision repair and strand-break repair pathways. APE1 hydrolyzes the phosphodiester bond at abasic sites, producing 5'-deoxyribose phosphate and the 3'-OH primer needed for repair synthesis. It also has additional repair activities, including the removal of 3'-blocking groups. APE1 is a powerful enzyme that absolutely requires Mg2+, but the stoichiometry and catalytic function of the divalent cation remain unresolved for APE1 and for other enzymes in the DNase I superfamily. Previously reported structures of DNA-free APE1 contained either Sm3+ or Pb2+ in the active site. However, these are poor surrogates for Mg2+ because Sm3+ is not a cofactor and Pb2+ inhibits APE1, and their coordination geometry is expected to differ from that of Mg2+. A crystal structure of human APE1 was solved at 1.92 Å resolution with a single Mg2+ ion in the active site. The structure reveals ideal octahedral coordination of Mg2+ via two carboxylate groups and four water molecules. One residue that coordinates Mg2+ directly and two that bind inner-sphere water molecules are strictly conserved in the DNase I superfamily. This structure, together with a recent structure of the enzyme-product complex, inform on the stoichiometry and the role of Mg2+ in APE1-catalyzed reactions.
- Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, 108 North Greene Street, Baltimore, MD 21201, USA.
Organizational Affiliation: