Anti-diabetic activity of insulin-degrading enzyme inhibitors mediated by multiple hormones.
Maianti, J.P., McFedries, A., Foda, Z.H., Kleiner, R.E., Du, X.Q., Leissring, M.A., Tang, W.J., Charron, M.J., Seeliger, M.A., Saghatelian, A., Liu, D.R.(2014) Nature 511: 94-98
- PubMed: 24847884 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13297
- Primary Citation Related Structures:
4LTE - PubMed Abstract:
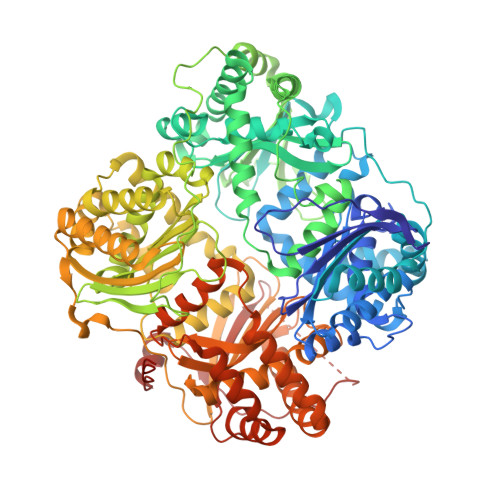
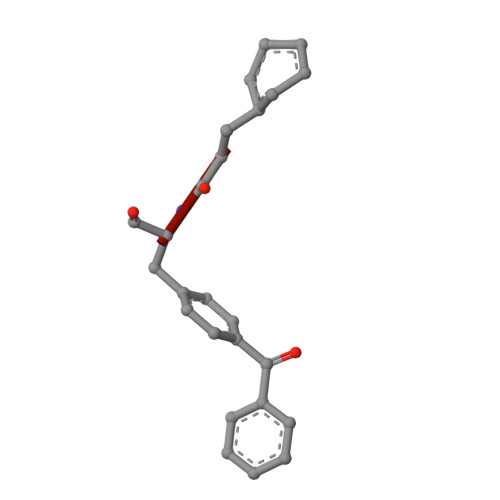
Despite decades of speculation that inhibiting endogenous insulin degradation might treat type-2 diabetes, and the identification of IDE (insulin-degrading enzyme) as a diabetes susceptibility gene, the relationship between the activity of the zinc metalloprotein IDE and glucose homeostasis remains unclear. Although Ide(-/-) mice have elevated insulin levels, they exhibit impaired, rather than improved, glucose tolerance that may arise from compensatory insulin signalling dysfunction. IDE inhibitors that are active in vivo are therefore needed to elucidate IDE's physiological roles and to determine its potential to serve as a target for the treatment of diabetes. Here we report the discovery of a physiologically active IDE inhibitor identified from a DNA-templated macrocycle library. An X-ray structure of the macrocycle bound to IDE reveals that it engages a binding pocket away from the catalytic site, which explains its remarkable selectivity. Treatment of lean and obese mice with this inhibitor shows that IDE regulates the abundance and signalling of glucagon and amylin, in addition to that of insulin. Under physiological conditions that augment insulin and amylin levels, such as oral glucose administration, acute IDE inhibition leads to substantially improved glucose tolerance and slower gastric emptying. These findings demonstrate the feasibility of modulating IDE activity as a new therapeutic strategy to treat type-2 diabetes and expand our understanding of the roles of IDE in glucose and hormone regulation.
- Department of Chemistry and Chemical Biology, Harvard University, 12 Oxford Street, Cambridge, Massachusetts 02138, USA.
Organizational Affiliation: