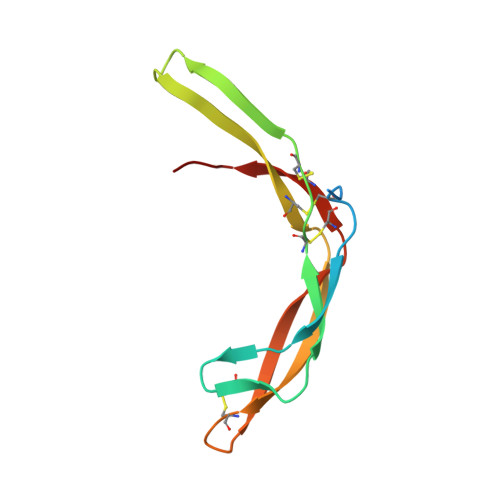
Highly reinforced structure of a C-terminal dimerization domain in von Willebrand factor.
Zhou, Y.F., Springer, T.A.(2014) Blood 123: 1785-1793
- PubMed: 24394662 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/blood-2013-11-523639
- Primary Citation Related Structures:
4NT5 - PubMed Abstract:
The C-terminal cystine knot (CK) (CTCK) domain in von Willebrand factor (VWF) mediates dimerization of proVWF in the endoplasmic reticulum and is essential for long multimers required for hemostatic function. The CTCK dimer crystal structure reveals highly elongated monomers with 2 β-ribbons and 4 intra-chain disulfides, including 3 in the CK. Dimerization buries an extensive interface of 1500 Å(2) corresponding to 32% of the surface of each monomer and forms a super β-sheet and 3 inter-chain disulfides. The shape, dimensions, and N-terminal connections of the crystal structure agree perfectly with previous electron microscopic images of VWF dimeric bouquets with the CTCK dimer forming a down-curved base. The dimer interface is suited to resist hydrodynamic force and disulfide reduction. CKs in each monomer flank the 3 inter-chain disulfides, and their presence in β-structures with dense backbone hydrogen bonds creates a rigid, highly crosslinked interface. The structure reveals the basis for von Willebrand disease phenotypes and the fold and disulfide linkages for CTCK domains in diverse protein families involved in barrier function, eye and inner ear development, insect coagulation and innate immunity, axon guidance, and signaling in extracellular matrices.
- Program in Cellular and Molecular Medicine, Children's Hospital Boston and Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA.
Organizational Affiliation: