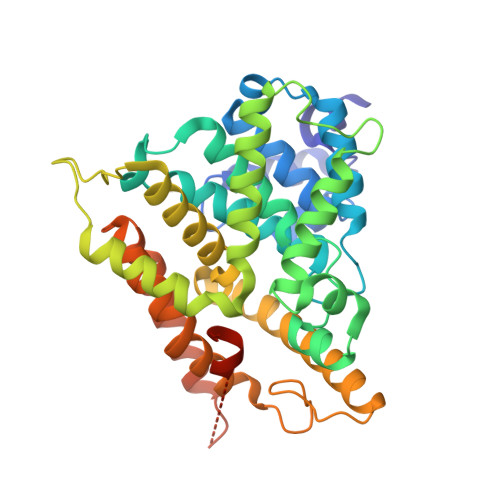
Discovery of triazines as selective PDE4B versus PDE4D inhibitors.
Hagen, T.J., Mo, X., Burgin, A.B., Fox, D., Zhang, Z., Gurney, M.E.(2014) Bioorg Med Chem Lett 24: 4031-4034
- PubMed: 24998378 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2014.06.002
- Primary Citation Related Structures:
4NW7 - PubMed Abstract:
In this study we report a series of triazine derivatives that are potent inhibitors of PDE4B. We also provide a series of structure activity relationships that demonstrate the triazine core can be used to generate subtype selective inhibitors of PDE4B versus PDE4D. A high resolution co-crystal structure shows that the inhibitors interact with a C-terminal regulatory helix (CR3) locking the enzyme in an inactive 'closed' conformation. The results show that the compounds interact with both catalytic domain and CR3 residues. This provides the first structure-based approach to engineer PDE4B-selective inhibitors.
- Department of Chemistry and Biochemistry, Northern Illinois University, DeKalb, IL 60115, USA. Electronic address: thagen@niu.edu.
Organizational Affiliation: