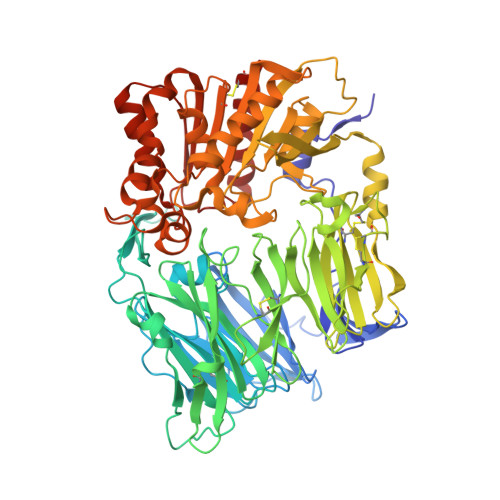
Omarigliptin (MK-3102): A Novel Long-Acting DPP-4 Inhibitor for Once-Weekly Treatment of Type 2 Diabetes.
Biftu, T., Sinha-Roy, R., Chen, P., Qian, X., Feng, D., Kuethe, J.T., Scapin, G., Gao, Y.D., Yan, Y., Krueger, D., Bak, A., Eiermann, G., He, J., Cox, J., Hicks, J., Lyons, K., He, H., Salituro, G., Tong, S., Patel, S., Doss, G., Petrov, A., Wu, J., Xu, S.S., Sewall, C., Zhang, X., Zhang, B., Thornberry, N.A., Weber, A.E.(2014) J Med Chem 57: 3205-3212
- PubMed: 24660890 Search on PubMed
- DOI: https://doi.org/10.1021/jm401992e
- Primary Citation Related Structures:
4PNZ - PubMed Abstract:
In our effort to discover DPP-4 inhibitors with added benefits over currently commercially available DPP-4 inhibitors, MK-3102 (omarigliptin), was identified as a potent and selective dipeptidyl peptidase 4 (DPP-4) inhibitor with an excellent pharmacokinetic profile amenable for once-weekly human dosing and selected as a clinical development candidate. This manuscript summarizes the mechanism of action, scientific rationale, medicinal chemistry, pharmacokinetic properties, and human efficacy data for omarigliptin, which is currently in phase 3 clinical development.
- Department of Discovery Chemistry, ‡Department of Metabolic Disorders, §Department of Pharmacology, ∥Department of Drug Metabolism, ⊥Basic Pharmaceutical Sciences, and #Department of Safety Assessment and Animal Resources, Merck & Co., Inc. , Whitehouse Station, New Jersey 08889, United States.
Organizational Affiliation: