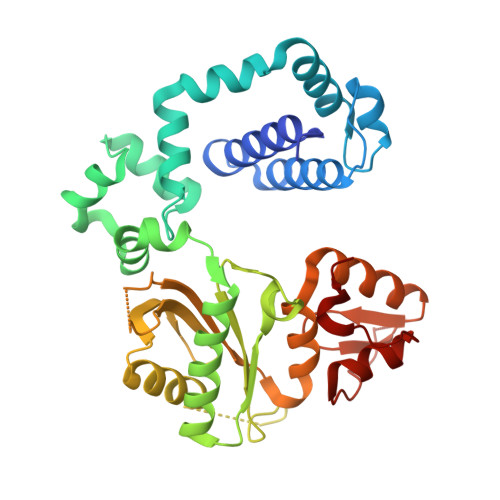
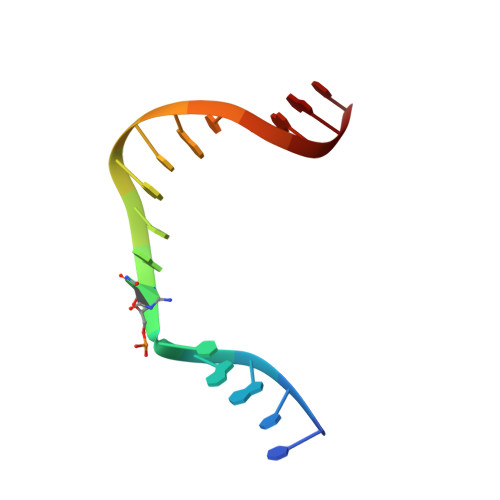
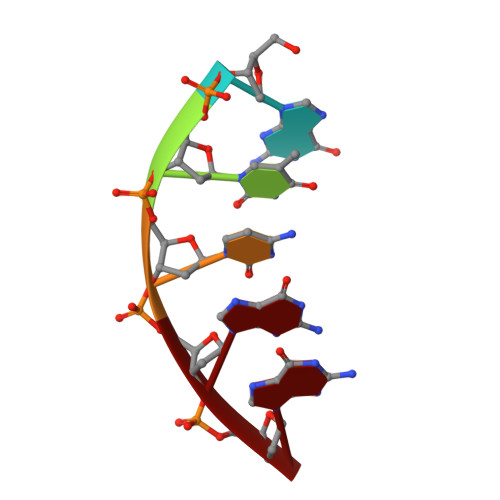
Crystal Structure of DNA Polymerase beta with DNA Containing the Base Lesion Spiroiminodihydantoin in a Templating Position.
Eckenroth, B.E., Fleming, A.M., Sweasy, J.B., Burrows, C.J., Doublie, S.(2014) Biochemistry 53: 2075-2077
- PubMed: 24649945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi500270e
- Primary Citation Related Structures:
4PPX - PubMed Abstract:
The first high-resolution crystal structure of spiroiminodihydantoin (dSp1) was obtained in the context of the DNA polymerase β active site and reveals two areas of significance. First, the structure verifies the recently determined S configuration at the spirocyclic carbon. Second, the distortion of the DNA duplex is similar to that of the single-oxidation product 8-oxoguanine. For both oxidized lesions, adaptation of the syn conformation results in similar backbone distortions in the DNA duplex. The resulting conformation positions the dSp1 A-ring as the base-pairing face whereas the B-ring of dSp1 protrudes into the major groove.
- Department of Microbiology and Molecular Genetics, University of Vermont , Stafford Hall, 95 Carrigan Drive, Burlington, Vermont 05405, United States.
Organizational Affiliation: