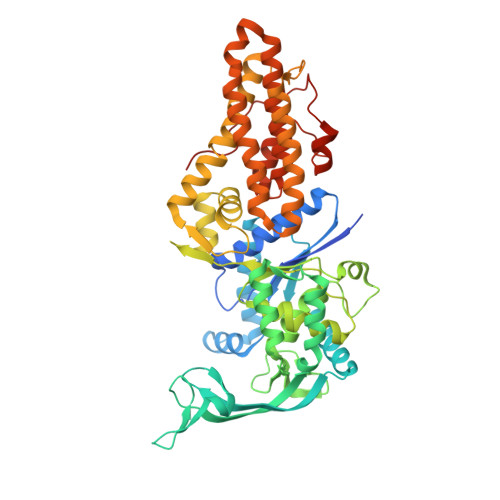
Brucella melitensis Methionyl-tRNA-Synthetase (MetRS), a Potential Drug Target for Brucellosis.
Ojo, K.K., Ranade, R.M., Zhang, Z., Dranow, D.M., Myers, J.B., Choi, R., Nakazawa Hewitt, S., Edwards, T.E., Davies, D.R., Lorimer, D., Boyle, S.M., Barrett, L.K., Buckner, F.S., Fan, E., Van Voorhis, W.C.(2016) PLoS One 11: e0160350-e0160350
- PubMed: 27500735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0160350
- Primary Citation Related Structures:
4DLP, 4PY2, 5K0S, 5K0T - PubMed Abstract:
We investigated Brucella melitensis methionyl-tRNA-synthetase (BmMetRS) with molecular, structural and phenotypic methods to learn if BmMetRS is a promising target for brucellosis drug development. Recombinant BmMetRS was expressed, purified from wild type Brucella melitensis biovar Abortus 2308 strain ATCC/CRP #DD-156 and screened by a thermal melt assay against a focused library of one hundred previously classified methionyl-tRNA-synthetase inhibitors of the blood stage form of Trypanosoma brucei. Three compounds showed appreciable shift of denaturation temperature and were selected for further studies on inhibition of the recombinant enzyme activity and cell viability against wild type B. melitensis strain 16M. BmMetRS protein complexed with these three inhibitors resolved into three-dimensional crystal structures and was analyzed. All three selected methionyl-tRNA-synthetase compounds inhibit recombinant BmMetRS enzymatic functions in an aminoacylation assay at varying concentrations. Furthermore, growth inhibition of B. melitensis strain 16M by the compounds was shown. Inhibitor-BmMetRS crystal structure models were used to illustrate the molecular basis of the enzyme inhibition. Our current data suggests that BmMetRS is a promising target for brucellosis drug development. However, further studies are needed to optimize lead compound potency, efficacy and safety as well as determine the pharmacokinetics, optimal dosage, and duration for effective treatment.
- Center for Emerging and Re-emerging Infectious Diseases (CERID), Department of Medicine, Division of Allergy and Infectious Diseases, University of Washington, Seattle, Washington, United States of America.
Organizational Affiliation: