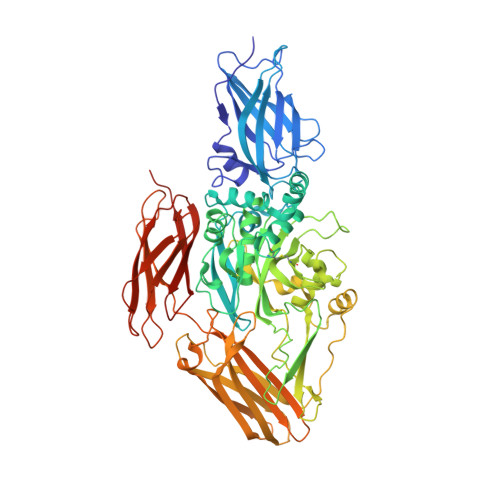
Crystal structure of transglutaminase 2 with GTP complex and amino acid sequence evidence of evolution of GTP binding site.
Jang, T.H., Lee, D.S., Choi, K., Jeong, E.M., Kim, I.G., Kim, Y.W., Chun, J.N., Jeon, J.H., Park, H.H.(2014) PLoS One 9: e107005-e107005
- PubMed: 25192068 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0107005
- Primary Citation Related Structures:
4PYG - PubMed Abstract:
Transglutaminase2 (TG2) is a multi-functional protein involved in various cellular processes, including apoptosis, differentiation, wound healing, and angiogenesis. The malfunction of TG2 causes many human disease including inflammatory disease, celiac disease, neurodegenerative diseases, tissue fibrosis, and cancers. Protein cross-linking activity, which is representative of TG2, is activated by calcium ions and suppressed by GTP. Here, we elucidated the structure of TG2 in complex with its endogenous inhibitor, GTP. Our structure showed why GTP is the optimal nucleotide for interacting with and inhibiting TG2. In addition, sequence comparison provided information describing the evolutionary scenario of GTP usage for controlling the activity of TG2.
- School of Biotechnology, Yeungnam University, Gyeongsan, South Korea; Graduate School of Biochemistry, Yeungnam University, Gyeongsan, South Korea.
Organizational Affiliation: