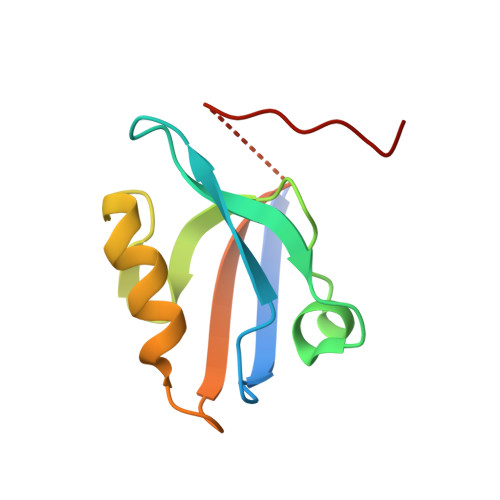
A structural portrait of the PDZ domain family.
Ernst, A., Appleton, B.A., Ivarsson, Y., Zhang, Y., Gfeller, D., Wiesmann, C., Sidhu, S.S.(2014) J Mol Biology 426: 3509-3519
- PubMed: 25158098 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2014.08.012
- Primary Citation Related Structures:
4Q2N, 4Q2O, 4Q2P, 4Q2Q - PubMed Abstract:
PDZ (PSD-95/Discs-large/ZO1) domains are interaction modules that typically bind to specific C-terminal sequences of partner proteins and assemble signaling complexes in multicellular organisms. We have analyzed the existing database of PDZ domain structures in the context of a specificity tree based on binding specificities defined by peptide-phage binding selections. We have identified 16 structures of PDZ domains in complex with high-affinity ligands and have elucidated four additional structures to assemble a structural database that covers most of the branches of the PDZ specificity tree. A detailed comparison of the structures reveals features that are responsible for the diverse specificities across the PDZ domain family. Specificity differences can be explained by differences in PDZ residues that are in contact with the peptide ligands, but these contacts involve both side-chain and main-chain interactions. Most PDZ domains bind peptides in a canonical conformation in which the ligand main chain adopts an extended β-strand conformation by interacting in an antiparallel fashion with a PDZ β-strand. However, a subset of PDZ domains bind peptides with a bent main-chain conformation and the specificities of these non-canonical domains could not be explained based on canonical structures. Our analysis provides a structural portrait of the PDZ domain family, which serves as a guide in understanding the structural basis for the diverse specificities across the family.
- Banting and Best Department of Medical Research and Department of Molecular Genetics, University of Toronto, The Donnelly Centre, 160 College Street, Toronto, ON M5S 3E1, Canada.
Organizational Affiliation: