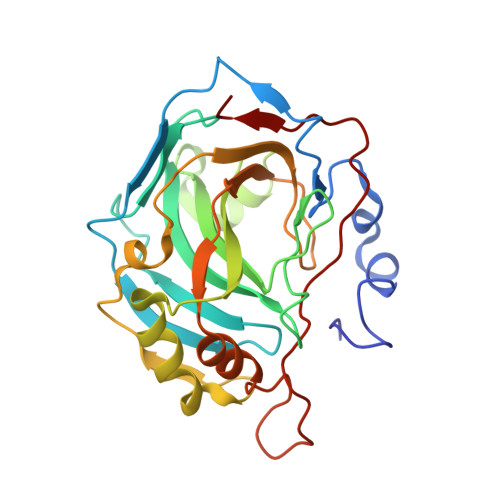
Out of the active site binding pocket for carbonic anhydrase inhibitors.
D'Ambrosio, K., Carradori, S., Monti, S.M., Buonanno, M., Secci, D., Vullo, D., Supuran, C.T., De Simone, G.(2014) Chem Commun (Camb) 51: 302-305
- PubMed: 25407638 Search on PubMed
- DOI: https://doi.org/10.1039/c4cc07320g
- Primary Citation Related Structures:
4QY3 - PubMed Abstract:
A structural study of the adduct which 2-benzylsulfinylbenzoic acid forms with human carbonic anhydrase II is reported, showing a binding mode completely different from any other class of carbonic anhydrase inhibitors investigated so far; this carboxylate binds in a pocket situated out of the enzyme active site.
- Istituto di Biostrutture e Bioimmagini-CNR, Via Mezzocannone 16, 80134, Naples, Italy. gdesimon@unina.it katia.dambrosio@cnr.it.
Organizational Affiliation: