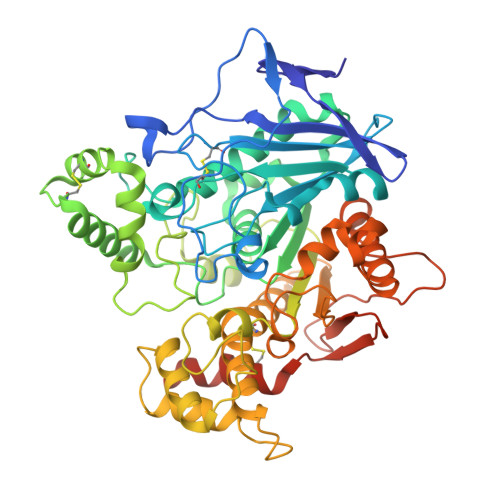
Structure-based development of nitroxoline derivatives as potential multifunctional anti-Alzheimer agents.
Knez, D., Brus, B., Coquelle, N., Sosic, I., Sink, R., Brazzolotto, X., Mravljak, J., Colletier, J.P., Gobec, S.(2015) Bioorg Med Chem 23: 4442-4452
- PubMed: 26116179 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2015.06.010
- Primary Citation Related Structures:
4XII - PubMed Abstract:
Tremendous efforts have been dedicated to the development of effective therapeutics against Alzheimer's disease, which represents the most common debilitating neurodegenerative disease. Multifunctional agents are molecules designed to have simultaneous effects on different pathological processes. Such compounds represent an emerging strategy for the development of effective treatments against Alzheimer's disease. Here, we report on the synthesis and biological evaluation of a series of nitroxoline-based analogs that were designed by merging the scaffold of 8-hydroxyquinoline with that of a known selective butyrylcholinesterase inhibitor that has promising anti-Alzheimer properties. Most strikingly, compound 8g inhibits self-induced aggregation of the amyloid beta peptide (Aβ1-42), inhibits with sub-micromolar potency butyrylcholinesterase (IC50=215 nM), and also selectively complexes Cu(2+). Our study thus designates this compound as a promising multifunctional agent for therapeutic treatment of Alzheimer's disease. The crystal structure of human butyrylcholinesterase in complex with compound 8g is also solved, which suggests ways to further optimize compounds featuring the 8-hydroxyquinoline scaffold.
- Faculty of Pharmacy, University of Ljubljana, Aškerčeva 7, 1000 Ljubljana, Slovenia.
Organizational Affiliation: