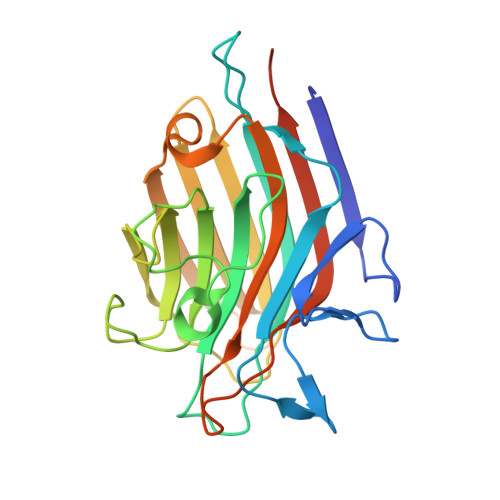
Crystal structure of a recombinant Vatairea macrocarpa seed lectin
Sousa, B.L., Silva-Filho, J.C., Kumar, P., Lyskowski, A., Bezerra, G.A., Delatorre, P., Rocha, B.A.M., Cunha, R.M.S., Nagano, C.S., Gruber, K., Cavada, B.S.(2016) Int J Biochem Cell Biol